microbiome
Microbe Normally Found in the Mouth May Drive Progression of Colorectal Cancer
Posted on by Dr. Monica M. Bertagnolli

Colorectal cancer is a leading cause of death from cancer in the United States. We know that risk of colorectal cancer goes up with age, certain coexisting health conditions, family history, smoking, alcohol use, and other factors. Researchers are also trying to learn more about what leads colorectal cancer to grow and spread. Now, findings from a new study supported in part by NIH add to evidence that colorectal tumor growth may be driven by a surprising bad actor: a microbe that’s normally found in the mouth.1
The findings, reported in Nature, suggest that a subtype of the bacterium Fusobacterium nucleatum has distinct genetic properties that may allow it to withstand acidic conditions in the stomach, infect colorectal tumors, and potentially drive their growth, which may lead to poorer patient outcomes. The discoveries suggest that the microbe could eventually be used as a target for detecting and treating colorectal cancer.
The study was conducted by a team led by Susan Bullman and Christopher D. Johnston at the Fred Hutchinson Cancer Center in Seattle. In 2022, the team published findings from a pair of studies implicating Fusobacterium nucleatum in the progression and spread of colorectal cancer.2,3 Their findings weren’t the first to suggest a link between the microbe and colorectal cancer. But their work offered important evidence that the microbe might alter colorectal tumors in ways that made them more likely to grow and spread. They also found that the microbe may affect the way colorectal cancer responds to or resists chemotherapy treatment.
Follow-up studies suggested there might be more to the story, pointing to the possibility that certain strains of the bacterium might differ from others in important ways. The findings suggested that there may be a more specific subtype, not yet defined, that was responsible for driving colorectal cancer growth.
To look deeper into this in the new study, Bullman and Johnston, with first author Martha Zepeda Rivera, analyzed a collection of 55 strains of the microbe taken from human colorectal cancer samples. They also compared these at the genetic level to another 80 strains of the microbe taken from the mouths of people who didn’t have cancer.
Their studies uncovered 483 genetic factors that turned up more often in Fusobacterium nucleatum from colorectal tumors. Those strains mainly belonged to a subspecies called Fusobacterium nucleatum animalis (Fna). More detailed study led to another surprise. The Fna included two genetically distinct groups or “clades” that had never been described, which the researchers called Fna C1 and C2. It turned out that only Fna C2 occurs at high levels in colorectal tumors.
The researchers found that this specific subtype within colorectal tumors carries 195 genetic factors that may allow it to grow more rapidly, withstand the acidic environment in the stomach, and take up residence in the gastrointestinal tract, where it can drive colorectal cancer growth. When the researchers infected a mouse model of colitis, a condition involving inflamed intestines that is a risk factor for colorectal cancer, they found that Fna C2 caused the development of more tumors compared to those infected with Fna C1.
Studies of tumors from 116 patients with colorectal cancer also showed more Fna C2. It was elevated in about 50% of cases. In fact, only this strain turned up more often in cancer compared to healthy tissue nearby. Stool samples of 627 people with colorectal cancer and 619 healthy people also showed more of this specific microbial strain in association with cancer.
This discovery is important because it suggests it’s only the Fna C2 subtype that’s associated with driving colorectal tumor growth, meaning it could help in the development of new methods for colorectal cancer screening and treatment. The researchers suggest it may one day even be possible to develop microbial-based therapies using modified versions of the bacterial strain to deliver treatments straight into tumors.
In addition, while the microbe is normally found in healthy mouths, it’s also enriched in periodontal (gum) disease, dental infections, and oral cancers.4 It will be interesting to learn more in future studies about the connections between various Fusobacterium nucleatum subtypes, oral health, and other health conditions throughout the body, including colorectal cancer.
References:
[1] Zepeda-Rivera M, et al. A distinct Fusobacterium nucleatum clade dominates the colorectal cancer niche. Nature. DOI: 10.1038/s41586-024-07182-w (2024).
[2] LaCourse KD, et al. The cancer chemotherapeutic 5-fluorouracil is a potent Fusobacterium nucleatum inhibitor and its activity is modified by intratumoral microbiota. Cell Rep. DOI: 10.1016/j.celrep.2022.111625 (2022).
[3] Galeano Niño JL, et al. Effect of the intratumoral microbiota on spatial and cellular heterogeneity in cancer. Nature. DOI: 10.1038/s41586-022-05435-0 (2022).
[4] Chen Y, et al. More Than Just a Periodontal Pathogen –the Research Progress on Fusobacterium nucleatum. Front Cell Infect Microbiol. DOI: 10.3389/fcimb.2022.815318 (2022).
NIH Support: National Institute of Dental and Craniofacial Research, National Cancer Institute
Changes in Human Microbiome Precede Alzheimer’s Cognitive Declines
Posted on by Lawrence Tabak, D.D.S., Ph.D.
In people with Alzheimer’s disease, the underlying changes in the brain associated with dementia typically begin many years—or even decades—before a diagnosis. While pinpointing the exact causes of Alzheimer’s remains a major research challenge, they likely involve a combination of genetic, environmental, and lifestyle factors. Now an NIH-funded study elucidates the role of another likely culprit that you may not have considered: the human gut microbiome, the trillions of diverse bacteria and other microbes that live primarily in our intestines [1].
Earlier studies had showed that the gut microbiomes of people with symptomatic Alzheimer’s disease differ from those of healthy people with normal cognition [2]. What this new work advances is that these differences arise early on in people who will develop Alzheimer’s, even before any obvious symptoms appear.
The science still has a ways to go before we’ll know if specific dietary changes can alter the gut microbiome and modify its influence on the brain in the right ways. But what’s exciting about this finding is it raises the possibility that doctors one day could test a patient’s stool sample to determine if what’s present from their gut microbiome correlates with greater early risk for Alzheimer’s dementia. Such a test would help doctors detect Alzheimer’s earlier and intervene sooner to slow or ideally even halt its advance.
The new findings, reported in the journal Science Translational Medicine, come from a research team led by Gautam Dantas and Beau Ances, Washington University School of Medicine, St. Louis. Ances is a clinician who treats and studies people with Alzheimer’s; Dantas is a basic researcher and expert on the gut microbiome.
The pair struck up a conversation one day about the possible connection between the gut microbiome and Alzheimer’s. While they knew about the earlier studies suggesting a link, they were surprised that nobody had looked at the gut microbiomes of people in the earliest, so-called preclinical, stages of the disease. That’s when dementia isn’t detectable, but the brain has formed amyloid-beta plaques, which are associated with Alzheimer’s.
To take a look, they enrolled 164 healthy volunteers, age 68 to 94, who performed normally on standard tests of cognition. They also collected stool samples from each volunteer and thoroughly analyzed them all the microbes from their gut microbiome. Study participants also kept food diaries and underwent extensive testing, including two types of brain scans, to look for signs of amyloid-beta plaques and tau protein accumulation that precede the onset of Alzheimer’s symptoms.
Among the volunteers, about a third (49 individuals) unfortunately had signs of early Alzheimer’s disease. And, as it turned out, their microbiomes showed differences, too.
The researchers found that those with preclinical Alzheimer’s disease had markedly different assemblages of gut bacteria. Their microbiomes differed in many of the bacterial species present. Those species-level differences also point to differences in the way their microbiomes would be expected to function at a metabolic level. These microbiome changes were observed even though the individuals didn’t seem to have any apparent differences in their diets.
The team also found that the microbiome changes correlated with amyloid-beta and tau levels in the brain. But they did not find any relationship to degenerative changes in the brain, which tend to happen later in people with Alzheimer’s.
The team is now conducting a five-year study that will follow volunteers to get a better handle on whether the differences observed in the gut microbiome are a cause or a consequence of the brain changes seen in Alzheimer’s. If it’s a cause, this discovery would raise the tantalizing possibility that specially formulated probiotics or fecal transplants that promote the growth of “good” bacteria over “bad” bacteria in the gut might slow the development of Alzheimer’s and its most devastating symptoms. It’s an exciting area of research and definitely one worth following in the years ahead.
References:
[1] Gut microbiome composition may be an indicator of preclinical Alzheimer’s disease. Ferreiro AL, Choi J, Ryou J, Newcomer EP, Thompson R, Bollinger RM, Hall-Moore C, Ndao IM, Sax L, Benzinger TLS, Stark SL, Holtzman DM, Fagan AM, Schindler SE, Cruchaga C, Butt OH, Morris JC, Tarr PI, Ances BM, Dantas G. Sci Transl Med. 2023 Jun 14;15(700):eabo2984. doi: 10.1126/scitranslmed.abo2984. Epub 2023 Jun 14. PMID: 37315112.
[2] Gut microbiome alterations in Alzheimer’s disease. Vogt NM, Kerby RL, Dill-McFarland KA, Harding SJ, Merluzzi AP, Johnson SC, Carlsson CM, Asthana S, Zetterberg H, Blennow K, Bendlin BB, Rey FE. Sci Rep. 2017 Oct 19;7(1):13537. doi: 10.1038/s41598-017-13601-y. PMID: 29051531; PMCID: PMC5648830.
Links:
Alzheimer’s Disease and Related Dementias (National Institute on Aging/NIH)
Video: How Alzheimer’s Changes the Brain (NIA)
Dantas Lab (Washington University School of Medicine. St. Louis)
Ances Bioimaging Laboratory (Washington University School of Medicine, St. Louis)
NIH Support: National Institute on Aging; National Institute of Diabetes and Digestive and Kidney Diseases
Unraveling the Role of the Skin Microbiome in Health and Disease
Posted on by Lindsey A. Criswell, M.D., M.P.H., D.Sc., National Institute of Arthritis and Musculoskeletal and Skin Diseases

Human skin is home to diverse ecosystems including bacteria, viruses, and fungi. These microbial communities comprise hundreds of species and are collectively known as the skin microbiome. The skin microbiome is thought to play a vital role in fending off disease-causing microorganisms (pathogens), boosting barrier protection, and aiding immune defenses.
Maintaining a balanced skin microbiome involves a complex and dynamic interplay among microorganisms, immune cells, skin cells, and other factors. In general, bacteria far outnumber viral, fungal, or other microbial species on the skin. Bacterial communities, which are strongly influenced by conditions such as skin moisture, temperature, and pH, vary widely across the body. For example, facial cheek skin hosts mostly Cutibacterium along with a bit of the skin fungus Malassezia. The heel is colonized by different types of bacteria including Staphylococcus and Corynebacteria.
In some diseases, such as acne and eczema, the skin microbiome is altered. Typically, this means an increase in pathogenic microorganisms and a decrease in beneficial ones. An altered skin microbiome can also be associated with inflammation, severe disease symptoms, and changes in the human immune system.
Heidi H. Kong is working to understand the role of the skin microbiome in health and disease. She is a senior investigator in the Intramural Research Program at NIH’s National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS) and an adjunct investigator at NIH’s National Cancer Institute (NCI).
More than a decade ago, Kong and Julie A. Segre, an intramural researcher at NIH’s National Human Genome Research Institute, analyzed the microbial makeup of healthy individuals. Kong swabbed the skin of these healthy volunteers in 20 different sites, from the forehead to the toenail. The study revealed that the surface of the human body provides various environmental niches, depending on whether the skin is moist, dry, or sebaceous (oily). Different bacterial species predominate in each niche. Kong and Segre were particularly interested in body areas that have predilections for disease. For example, psoriasis is often found on the outside of elbows and knees, and the back of the scalp.
Earlier this year, Kong and Segre published another broad analysis of the human skin microbiome [1] in collaboration with scientists at the European Molecular Biology Laboratory, European Bioinformatics Institute (EMBL-EBI), United Kingdom. This new catalog, called the Skin Microbial Genome Collection, is thought to identify about 85 percent of the microorganisms present on healthy skin from 19 body sites. It documents more than 600 bacterial species—including 174 that were discovered during the study—as well as more than 6,900 viruses and some fungi, including three newly discovered species.
Kong’s work has provided compelling evidence that the human immune system plays a role in shaping the skin microbiome. In 2018, she, Segre, and colleagues from the intramural programs of NCI and NIH’s National Institute of Allergy and Infectious Diseases analyzed skin from eight different sites on 27 people with a rare primary immunodeficiency disease known as DOCK8 deficiency [2].
People with the condition have recurrent infections in the skin, sinuses, and airways, and are susceptible to different cancers. Kong and colleagues found that the skin of people with DOCK8 deficiency contains significantly more DNA viruses (90 percent of the skin microbiome on average) than people without the condition (6 or 7 percent of the skin microbiome).
Other researchers are hoping to leverage features of the microbiome to develop targeted therapies for skin diseases. Richard L. Gallo, a NIAMS grantee at the University of California, San Diego, is currently focused on acne and eczema (also called atopic dermatitis). Acne is associated with certain strains of Cutibacterium acnes (C. acnes, formerly called Propionibacterium acnes or P. acnes). Eczema is often associated with Staphylococcus aureus (S. aureus).
Severe cases of acne and eczema are commonly treated with broad-spectrum antibiotics, which wipe out most of the bacteria, including beneficial species. The goal of microbiome-targeted therapy is to kill only the disease-associated bacteria and avoid increasing the risk that some strains will develop antibiotic resistance.
In 2020, Gallo and colleagues identified a strain of Staphylococcus capitis from healthy human skin (S. capitis E12) that selectively inhibits the growth of C. acnes without negatively impacting other bacteria or human skin cells [3]. S. capitis E12 produces four different toxins that act together to target C. acnes. The research team created an extract of the four toxins and tested it using animal models. In most cases, the extract was more potent at killing C. acnes—including acne-associated strains—than several commonly prescribed antibiotics (erythromycin, tetracycline, and clindamycin). And, unlike antibiotics, the extract does not appear to promote drug-resistance, at least for the 20 generations observed by the researchers.
Eczema is a chronic, relapsing disease characterized by skin that is dry, itchy, inflamed, and prone to infection, including by pathogens such as S. aureus and herpes virus. Although the cause of eczema is unknown, the condition is associated with human genetic mutations, disruption of the skin’s barrier, inflammation-triggering allergens, and imbalances in the skin microbiome.
In 2017, Gallo’s research team discovered that, in healthy human skin, certain strains of Staphylococcus hominis and Staphylococcus epidermis produce potent antimicrobial molecules known as lantibiotics [4]. These beneficial strains are far less common on the skin of people with eczema. The lantibiotics work synergistically with LL-37, an antimicrobial molecule produced by the human immune system, to selectively kill S. aureus, including methicillin-resistant strains (MRSA).
Gallo and his colleagues then examined the safety and therapeutic potential of these beneficial strains isolated from the human skin microbiome. In animal tests, strains of S. hominis and S. epidermis that produce lantibiotics killed S. aureus and blocked production of its toxin.
Gallo’s group has now expanded their work to early studies in humans. In 2021, two independent phase 1 clinical trials [5,6] conducted by Gallo and his colleagues investigated the effects of these strains on people with eczema. These double-blind, placebo-controlled trials involved one-week of topical application of beneficial bacteria to the forearm of adults with S. aureus-positive eczema. The results demonstrated that the treatment was safe, showed a significant decrease in S. aureus, and improved eczema symptoms in most patients. This is encouraging news for those hoping to develop microbiome-targeted therapy for inflammatory skin diseases.
As research on the skin microbiome advances on different fronts, it will provide deeper insight into the multi-faceted microbial communities that are so critical to health and disease. One day, we may even be able to harness the microbiome as a source of therapeutics to alleviate inflammation, promote wound healing, or suppress certain skin cancers.
References:
[1] Integrating cultivation and metagenomics for a multi-kingdom view of skin microbiome diversity and functions. Saheb Kashaf S, Proctor DM, Deming C, Saary P, Hölzer M; NISC Comparative Sequencing Program, Taylor ME, Kong HH, Segre JA, Almeida A, Finn RD. Nat Microbiol. 2022 Jan;7(1):169-179.
[2] Expanded skin virome in DOCK8-deficient patients. Tirosh O, Conlan S, Deming C, Lee-Lin SQ, Huang X; NISC Comparative Sequencing Program, Su HC, Freeman AF, Segre JA, Kong HH. Nat Med. 2018 Dec;24(12):1815-1821.
[3] Identification of a human skin commensal bacterium that selectively kills Cutibacterium acnes. O’Neill AM, Nakatsuji T, Hayachi A, Williams MR, Mills RH, Gonzalez DJ, Gallo RL. J Invest Dermatol. 2020 Aug;140(8):1619-1628.e2.
[4] Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Nakatsuji T, Chen TH, Narala S, Chun KA, Two AM, Yun T, Shafiq F, Kotol PF, Bouslimani A, Melnik AV, Latif H, Kim JN, Lockhart A, Artis K, David G, Taylor P, Streib J, Dorrestein PC, Grier A, Gill SR, Zengler K, Hata TR, Leung DY, Gallo RL. Sci Transl Med. 2017 Feb 22;9(378):eaah4680.
[5] Development of a human skin commensal microbe for bacteriotherapy of atopic dermatitis and use in a phase 1 randomized clinical trial. Nakatsuji T, Hata TR, Tong Y, Cheng JY, Shafiq F, Butcher AM, Salem SS, Brinton SL, Rudman Spergel AK, Johnson K, Jepson B, Calatroni A, David G, Ramirez-Gama M, Taylor P, Leung DYM, Gallo RL. Nat Med. 2021 Apr;27(4):700-709.
[6] Use of autologous bacteriotherapy to treat Staphylococcus aureus in patients with atopic dermatitis: A randomized double-blind clinical trial. Nakatsuji T, Gallo RL, Shafiq F, Tong Y, Chun K, Butcher AM, Cheng JY, Hata TR. JAMA Dermatol. 2021 Jun 16;157(8):978-82.
Links:
Acne (National Institute of Arthritis and Musculoskeletal and Skin Diseases/NIH)
Atopic Dermatitis (NIAMS)
Cutaneous Microbiome and Inflammation Laboratory, Heidi Kong (NIAMS)
Julie Segre (National Human Genome Research Institute/NIH)
Gallo Lab (University of California, San Diego)
[Note: Acting NIH Director Lawrence Tabak has asked the heads of NIH’s Institutes and Centers (ICs) to contribute occasional guest posts to the blog to highlight some of the cool science that they support and conduct. This is the fifth in the series of NIH IC guest posts that will run until a new permanent NIH director is in place.]
Celebrating NIH Science, Blogs, and Blog Readers!
Posted on by Dr. Francis Collins
Happy holidays to one and all! As you may have heard, this is my last holiday season as the Director of the National Institutes of Health (NIH)—a post that I’ve held for the past 12 years and four months under three U.S. Presidents. And, wow, it really does seem like only yesterday that I started this blog!
At the blog’s outset, I said my goal was to “highlight new discoveries in biology and medicine that I think are game changers, noteworthy, or just plain cool.” More than 1,100 posts, 10 million unique visitors, and 13.7 million views later, I hope you’ll agree that goal has been achieved. I’ve also found blogging to be a whole lot of fun, as well as a great way to expand my own horizons and share a little of what I’ve learned about biomedical advances with people all across the nation and around the world.
So, as I sign off as NIH Director and return to my lab at NIH’s National Human Genome Research Institute (NHGRI), I want to thank everyone who’s ever visited this Blog—from high school students to people with health concerns, from biomedical researchers to policymakers. I hope that the evidence-based information that I’ve provided has helped and informed my readers in some small way.
In this my final post, I’m sharing a short video (see above) that highlights just a few of the blog’s many spectacular images, many of them produced by NIH-funded scientists during the course of their research. In the video, you’ll see a somewhat quirky collection of entries, but hopefully you will sense my enthusiasm for the potential of biomedical research to fight human disease and improve human health—from innovative immunotherapies for treating cancer to the gift of mRNA vaccines to combat a pandemic.
Over the years, I’ve blogged about many of the bold, new frontiers of biomedicine that are now being explored by research teams supported by NIH. Who would have imagined that, within the span of a dozen years, precision medicine would go from being an interesting idea to a driving force behind the largest-ever NIH cohort seeking to individualize the prevention and treatment of common disease? Or that today we’d be deep into investigations of precisely how the human brain works, as well as how human health may benefit from some of the trillions of microbes that call our bodies home?
My posts also delved into some of the amazing technological advances that are enabling breakthroughs across a wide range of scientific fields. These innovative technologies include powerful new ways of mapping the atomic structures of proteins, editing genetic material, and designing improved gene therapies.
So, what’s next for NIH? Let me assure you that NIH is in very steady hands as it heads into a bright horizon brimming with exceptional opportunities for biomedical research. Like you, I look forward to discoveries that will lead us even closer to the life-saving answers that we all want and need.
While we wait for the President to identify a new NIH director, Lawrence Tabak, who has been NIH’s Principal Deputy Director and my right arm for the last decade, will serve as Acting NIH Director. So, keep an eye out for his first post in early January!
As for me, I’ll probably take a little time to catch up on some much-needed sleep, do some reading and writing, and hopefully get out for a few more rides on my Harley with my wife Diane. But there’s plenty of work to do in my lab, where the focus is on type 2 diabetes and a rare disease of premature aging called Hutchinson-Gilford Progeria Syndrome. I’m excited to pursue those research opportunities and see where they lead.
In closing, I’d like to extend my sincere thanks to each of you for your interest in hearing from the NIH Director—and supporting NIH research—over the past 12 years. It’s been an incredible honor to serve you at the helm of this great agency that’s often called the National Institutes of Hope. And now, for one last time, Diane and I take great pleasure in sending you and your loved ones our most heartfelt wishes for Happy Holidays and a Healthy New Year!
Could A Gut-Brain Connection Help Explain Autism?
Posted on by Dr. Francis Collins

You might think nutrient-sensing cells in the human gastrointestinal (GI) tract would have no connection whatsoever to autism spectrum disorder (ASD). But if Diego Bohórquez’s “big idea” is correct, these GI cells, called neuropods, could one day help to provide a direct link into understanding and treating some aspects of autism and other brain disorders.
Bohórquez, a researcher at Duke University, Durham, NC, recently discovered that cells in the intestine, previously known for their hormone-releasing ability, form extensions similar to neurons. He also found that those extensions connect to nerve fibers in the gut, which relay signals to the vagus nerve and onward to the brain. In fact, he found that those signals reach the brain in milliseconds [1].
Bohórquez has dedicated his lab to studying this direct, high-speed hookup between gut and brain and its impact on nutrient sensing, eating, and other essential behaviors. Now, with support from a 2019 NIH Director’s New Innovator Award, he will also explore the potential for treating autism and other brain disorders with drugs that act on the gut.
Bohórquez became interested in autism and its possible link to the gut-brain connection after a chance encounter with Geraldine Dawson, director of the Duke Center for Autism and Brain Development. Dawson mentioned that autism typically affects multiple organ systems.
With further reading, he discovered that kids with autism frequently cope with GI issues, including bowel inflammation, abdominal pain, constipation, and/or diarrhea [2]. They often also show unusual food-related behaviors, such as being extremely picky eaters. But his curiosity was especially piqued by evidence that certain gut microbes can influence abnormal behaviors in mice that model autism.
With his New Innovator Award, Bohórquez will study neuropods and the gut-brain connection in a mouse model of autism. Using the tools of optogenetics, which make it possible to activate cells with light, he’ll also see whether autism-like symptoms in mice can be altered or alleviated by controlling neuropods in the gut. Those symptoms include anxiety, repetitive behaviors, and lack of interest in interacting with other mice. He’ll also explore changes in the animals’ eating habits.
In another line of study, he will take advantage of intestinal tissue samples collected from people with autism. He’ll use those tissues to grow and then examine miniature intestinal “organoids,” looking for possible evidence that those from people with autism are different from others.
For the millions of people now living with autism, no truly effective drug therapies are available to help to manage the condition and its many behavioral and bodily symptoms. Bohórquez hopes one day to change that with drugs that act safely on the gut. In the meantime, he and his fellow “GASTRONAUTS” look forward to making some important and fascinating discoveries in the relatively uncharted territory where the gut meets the brain.
References:
[1] A gut-brain neural circuit for nutrient sensory transduction. Kaelberer MM, Buchanan KL, Klein ME, Barth BB, Montoya MM, Shen X, Bohórquez DV. Science. 2018 Sep 21;361(6408).
[2] Association of maternal report of infant and toddler gastrointestinal symptoms with autism: evidence from a prospective birth cohort. Bresnahan M, Hornig M, Schultz AF, Gunnes N, Hirtz D, Lie KK, Magnus P, Reichborn-Kjennerud T, Roth C, Schjølberg S, Stoltenberg C, Surén P, Susser E, Lipkin WI. JAMA Psychiatry. 2015 May;72(5):466-474.
Links:
Autism Spectrum Disorder (National Institute of Mental Health/NIH)
Bohórquez Lab (Duke University, Durham, NC)
Bohórquez Project Information (NIH RePORTER)
NIH Director’s New Innovator Award (Common Fund)
NIH Support: Common Fund; National Institute of Mental Health
Celebrating 2019 Biomedical Breakthroughs
Posted on by Dr. Francis Collins

Happy New Year! As we say goodbye to the Teens, let’s take a look back at 2019 and some of the groundbreaking scientific discoveries that closed out this remarkable decade.
Each December, the reporters and editors at the journal Science select their breakthrough of the year, and the choice for 2019 is nothing less than spectacular: An international network of radio astronomers published the first image of a black hole, the long-theorized cosmic singularity where gravity is so strong that even light cannot escape [1]. This one resides in a galaxy 53 million light-years from Earth! (A light-year equals about 6 trillion miles.)
Though the competition was certainly stiff in 2019, the biomedical sciences were well represented among Science’s “runner-up” breakthroughs. They include three breakthroughs that have received NIH support. Let’s take a look at them:
In a first, drug treats most cases of cystic fibrosis: Last October, two international research teams reported the results from phase 3 clinical trials of the triple drug therapy Trikafta to treat cystic fibrosis (CF). Their data showed Trikafta effectively compensates for the effects of a mutation carried by about 90 percent of people born with CF. Upon reviewing these impressive data, the Food and Drug Administration (FDA) approved Trikafta, developed by Vertex Pharmaceuticals.
The approval of Trikafta was a wonderful day for me personally, having co-led the team that isolated the CF gene 30 years ago. A few years later, I wrote a song called “Dare to Dream” imagining that wonderful day when “the story of CF is history.” Though we’ve still got more work to do, we’re getting a lot closer to making that dream come true. Indeed, with the approval of Trikafta, most people with CF have for the first time ever a real chance at managing this genetic disease as a chronic condition over the course of their lives. That’s a tremendous accomplishment considering that few with CF lived beyond their teens as recently as the 1980s.
Such progress has been made possible by decades of work involving a vast number of researchers, many funded by NIH, as well as by more than two decades of visionary and collaborative efforts between the Cystic Fibrosis Foundation and Aurora Biosciences (now, Vertex) that built upon that fundamental knowledge of the responsible gene and its protein product. Not only did this innovative approach serve to accelerate the development of therapies for CF, it established a model that may inform efforts to develop therapies for other rare genetic diseases.
Hope for Ebola patients, at last: It was just six years ago that news of a major Ebola outbreak in West Africa sounded a global health emergency of the highest order. Ebola virus disease was then recognized as an untreatable, rapidly fatal illness for the majority of those who contracted it. Though international control efforts ultimately contained the spread of the virus in West Africa within about two years, over 28,600 cases had been confirmed leading to more than 11,000 deaths—marking the largest known Ebola outbreak in human history. Most recently, another major outbreak continues to wreak havoc in northeastern Democratic Republic of Congo (DRC), where violent civil unrest is greatly challenging public health control efforts.
As troubling as this news remains, 2019 brought a needed breakthrough for the millions of people living in areas susceptible to Ebola outbreaks. A randomized clinical trial in the DRC evaluated four different drugs for treating acutely infected individuals, including an antibody against the virus called mAb114, and a cocktail of anti-Ebola antibodies referred to as REGN-EB3. The trial’s preliminary data showed that about 70 percent of the patients who received either mAb114 or the REGN-EB3 antibody cocktail survived, compared with about half of those given either of the other two medicines.
So compelling were these preliminary results that the trial, co-sponsored by NIH’s National Institute of Allergy and Infectious Diseases (NIAID) and the DRC’s National Institute for Biomedical Research, was halted last August. The results were also promptly made public to help save lives and stem the latest outbreak. All Ebola patients in the DRC treatment centers now are treated with one or the other of these two options. The trial results were recently published.
The NIH-developed mAb114 antibody and the REGN-EB3 cocktail are the first therapeutics to be shown in a scientifically rigorous study to be effective at treating Ebola. This work also demonstrates that ethically sound clinical research can be conducted under difficult conditions in the midst of a disease outbreak. In fact, the halted study was named Pamoja Tulinde Maisha (PALM), which means “together save lives” in Kiswahili.
To top off the life-saving progress in 2019, the FDA just approved the first vaccine for Ebola. Called Ervebo (earlier rVSV-ZEBOV), this single-dose injectable vaccine is a non-infectious version of an animal virus that has been genetically engineered to carry a segment of a gene from the Zaire species of the Ebola virus—the virus responsible for the current DRC outbreak and the West Africa outbreak. Because the vaccine does not contain the whole Zaire virus, it can’t cause Ebola. Results from a large study in Guinea conducted by the WHO indicated that the vaccine offered substantial protection against Ebola virus disease. Ervebo, produced by Merck, has already been given to over 259,000 individuals as part of the response to the DRC outbreak. The NIH has supported numerous clinical trials of the vaccine, including an ongoing study in West Africa.
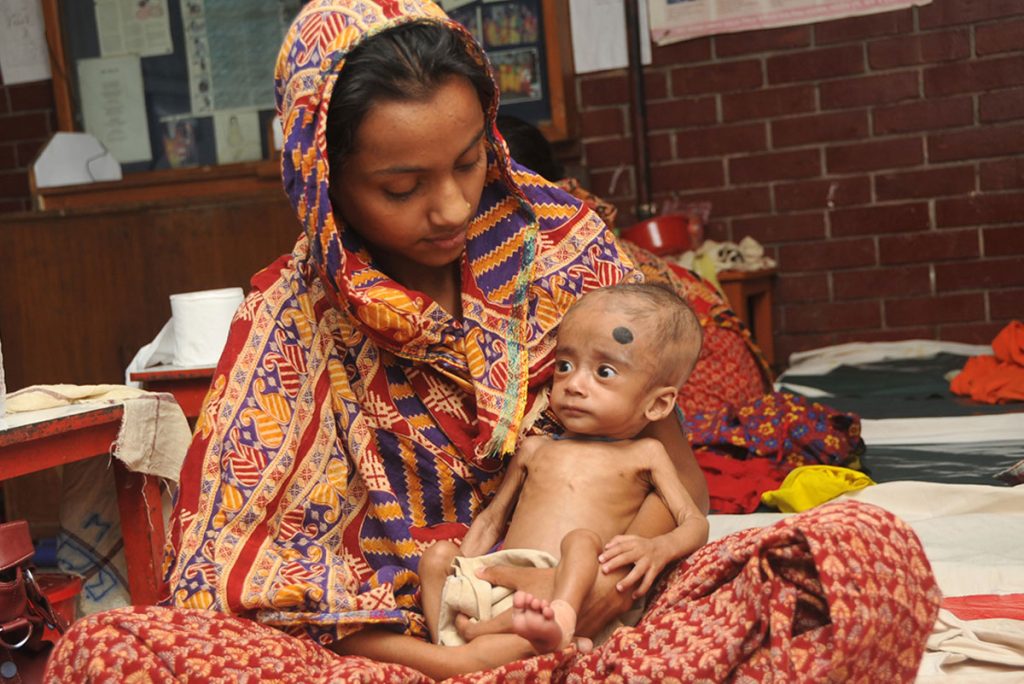
Microbes combat malnourishment: Researchers discovered a few years ago that abnormal microbial communities, or microbiomes, in the intestine appear to contribute to childhood malnutrition. An NIH-supported research team followed up on this lead with a study of kids in Bangladesh, and it published last July its groundbreaking finding: that foods formulated to repair the “gut microbiome” helped malnourished kids rebuild their health. The researchers were able to identify a network of 15 bacterial species that consistently interact in the gut microbiomes of Bangladeshi children. In this month-long study, this bacterial network helped the researchers characterize a child’s microbiome and/or its relative state of repair.
But a month isn’t long enough to determine how the new foods would help children grow and recover. The researchers are conducting a similar study that is much longer and larger. Globally, malnutrition affects an estimated 238 million children under the age 5, stunting their normal growth, compromising their health, and limiting their mental development. The hope is that these new foods and others adapted for use around the world soon will help many more kids grow up to be healthy adults.
Measles Resurgent: The staff at Science also listed their less-encouraging 2019 Breakdowns of the Year, and unfortunately the biomedical sciences made the cut with the return of measles in the U.S. Prior to 1963, when the measles vaccine was developed, 3 to 4 million Americans were sickened by measles each year. Each year about 500 children would die from measles, and many more would suffer lifelong complications. As more people were vaccinated, the incidence of measles plummeted. By the year 2000, the disease was even declared eliminated from the U.S.
But, as more parents have chosen not to vaccinate their children, driven by the now debunked claim that vaccines are connected to autism, measles has made a very preventable comeback. Last October, the Centers for Disease Control and Prevention (CDC) reported an estimated 1,250 measles cases in the United States at that point in 2019, surpassing the total number of cases reported annually in each of the past 25 years.
The good news is those numbers can be reduced if more people get the vaccine, which has been shown repeatedly in many large and rigorous studies to be safe and effective. The CDC recommends that children should receive their first dose by 12 to 15 months of age and a second dose between the ages of 4 and 6. Older people who’ve been vaccinated or have had the measles previously should consider being re-vaccinated, especially if they live in places with low vaccination rates or will be traveling to countries where measles are endemic.
Despite this public health breakdown, 2019 closed out a memorable decade of scientific discovery. The Twenties will build on discoveries made during the Teens and bring us even closer to an era of precision medicine to improve the lives of millions of Americans. So, onward to 2020—and happy New Year!
Reference:
[1] 2019 Breakthrough of the Year. Science, December 19, 2019.
NIH Support: These breakthroughs represent the culmination of years of research involving many investigators and the support of multiple NIH institutes.
Gut-Dwelling Bacterium Consumes Parkinson’s Drug
Posted on by Dr. Francis Collins

Scientists continue to uncover the many fascinating ways in which the trillions of microbes that inhabit the human body influence our health. Now comes yet another surprising discovery: a medicine-eating bacterium residing in the human gut that may affect how well someone responds to the most commonly prescribed drug for Parkinson’s disease.
There have been previous hints that gut microbes might influence the effectiveness of levodopa (L-dopa), which helps to ease the stiffness, rigidity, and slowness of movement associated with Parkinson’s disease. Now, in findings published in Science, an NIH-funded team has identified a specific, gut-dwelling bacterium that consumes L-dopa [1]. The scientists have also identified the bacterial genes and enzymes involved in the process.
Parkinson’s disease is a progressive neurodegenerative condition in which the dopamine-producing cells in a portion of the brain called the substantia nigra begin to sicken and die. Because these cells and their dopamine are critical for controlling movement, their death leads to the familiar tremor, difficulty moving, and the characteristic slow gait. As the disease progresses, cognitive and behavioral problems can take hold, including depression, personality shifts, and sleep disturbances.
For the 10 million people in the world now living with this neurodegenerative disorder, and for those who’ve gone before them, L-dopa has been for the last 50 years the mainstay of treatment to help alleviate those motor symptoms. The drug is a precursor of dopamine, and, unlike dopamine, it has the advantage of crossing the blood-brain barrier. Once inside the brain, an enzyme called DOPA decarboxylase converts L-dopa to dopamine.
Unfortunately, only a small fraction of L-dopa ever reaches the brain, contributing to big differences in the drug’s efficacy from person to person. Since the 1970s, researchers have suspected that these differences could be traced, in part, to microbes in the gut breaking down L-dopa before it gets to the brain.
To take a closer look in the new study, Vayu Maini Rekdal and Emily Balskus, Harvard University, Cambridge, MA, turned to data from the NIH-supported Human Microbiome Project (HMP). The project used DNA sequencing to identify and characterize the diverse collection of microbes that populate the healthy human body.
The researchers sifted through the HMP database for bacterial DNA sequences that appeared to encode an enzyme capable of converting L-dopa to dopamine. They found what they were looking for in a bacterial group known as Enterococcus, which often inhabits the human gastrointestinal tract.
Next, they tested the ability of seven representative Enterococcus strains to transform L-dopa. Only one fit the bill: a bacterium called Enterococcus faecalis, which commonly resides in a healthy gut microbiome. In their tests, this bacterium avidly consumed all the L-dopa, using its own version of a decarboxylase enzyme. When a specific gene in its genome was inactivated, E. faecalis stopped breaking down L-dopa.
These studies also revealed variability among human microbiome samples. In seven stool samples, the microbes tested didn’t consume L-dopa at all. But in 12 other samples, microbes consumed 25 to 98 percent of the L-dopa!
The researchers went on to find a strong association between the degree of L-dopa consumption and the abundance of E. faecalis in a particular microbiome sample. They also showed that adding E. faecalis to a sample that couldn’t consume L-dopa transformed it into one that could.
So how can this information be used to help people with Parkinson’s disease? Answers are already appearing. The researchers have found a small molecule that prevents the E. faecalis decarboxylase from modifying L-dopa—without harming the microbe and possibly destabilizing an otherwise healthy gut microbiome.
The finding suggests that the human gut microbiome might hold a key to predicting how well people with Parkinson’s disease will respond to L-dopa, and ultimately improving treatment outcomes. The finding also serves to remind us just how much the microbiome still has to tell us about human health and well-being.
Reference:
[1] Discovery and inhibition of an interspecies gut bacterial pathway for Levodopa metabolism. Maini Rekdal V, Bess EN, Bisanz JE, Turnbaugh PJ, Balskus EP. Science. 2019 Jun 14;364(6445).
Links:
Parkinson’s Disease Information Page (National Institute of Neurological Disorders and Stroke/NIH)
Balskus Lab (Harvard University, Cambridge, MA)
NIH Support: National Institute of General Medical Sciences; National Heart, Lung, and Blood Institute
Next Page