infants
Studies Confirm COVID-19 mRNA Vaccines Safe, Effective for Pregnant Women
Posted on by Dr. Francis Collins
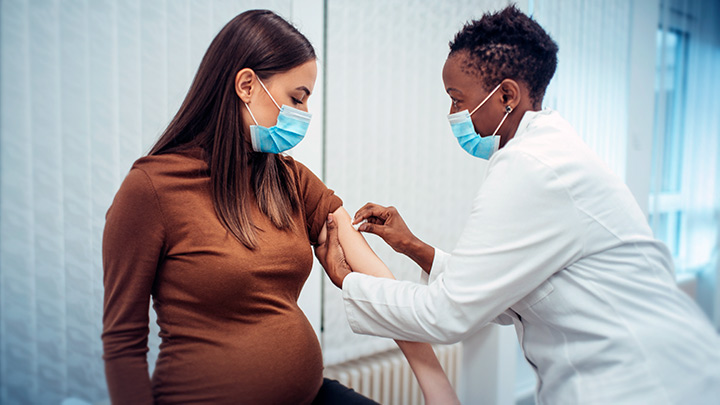
Clinical trials have shown that COVID-19 vaccines are remarkably effective in protecting those age 12 and up against infection by the coronavirus SARS-CoV-2. The expectation was that they would work just as well to protect pregnant women. But because pregnant women were excluded from the initial clinical trials, hard data on their safety and efficacy in this important group has been limited.
So, I’m pleased to report results from two new studies showing that the two COVID-19 mRNA vaccines now available in the United States appear to be completely safe for pregnant women. The women had good responses to the vaccines, producing needed levels of neutralizing antibodies and immune cells known as memory T cells, which may offer more lasting protection. The research also indicates that the vaccines might offer protection to infants born to vaccinated mothers.
In one study, published in JAMA [1], an NIH-supported team led by Dan Barouch, Beth Israel Deaconess Medical Center and Harvard Medical School, Boston, wanted to learn whether vaccines would protect mother and baby. To find out, they enrolled 103 women, aged 18 to 45, who chose to get either the Pfizer/BioNTech or Moderna mRNA vaccines from December 2020 through March 2021.
The sample included 30 pregnant women,16 women who were breastfeeding, and 57 women who were neither pregnant nor breastfeeding. Pregnant women in the study got their first dose of vaccine during any trimester, although most got their shots in the second or third trimester. Overall, the vaccine was well tolerated, although some women in each group developed a transient fever after the second vaccine dose, a common side effect in all groups that have been studied.
After vaccination, women in all groups produced antibodies against SARS-CoV-2. Importantly, those antibodies neutralized SARS-CoV-2 variants of concern. The researchers also found those antibodies in infant cord blood and breast milk, suggesting that they were passed on to afford some protection to infants early in life.
The other NIH-supported study, published in the journal Obstetrics & Gynecology, was conducted by a team led by Jeffery Goldstein, Northwestern’s Feinberg School of Medicine, Chicago [2]. To explore any possible safety concerns for pregnant women, the team took a first look for any negative effects of vaccination on the placenta, the vital organ that sustains the fetus during gestation.
The researchers detected no signs that the vaccines led to any unexpected damage to the placenta in this study, which included 84 women who received COVID-19 mRNA vaccines during pregnancy, most in the third trimester. As in the other study, the team found that vaccinated pregnant women showed a robust response to the vaccine, producing needed levels of neutralizing antibodies.
Overall, both studies show that COVID-19 mRNA vaccines are safe and effective in pregnancy, with the potential to benefit both mother and baby. Pregnant women also are more likely than women who aren’t pregnant to become severely ill should they become infected with this devastating coronavirus [3]. While pregnant women are urged to consult with their obstetrician about vaccination, growing evidence suggests that the best way for women during pregnancy or while breastfeeding to protect themselves and their families against COVID-19 is to roll up their sleeves and get either one of the mRNA vaccines now authorized for emergency use.
References:
[1] Immunogenicity of COVID-19 mRNA vaccines in pregnant and lactating women. Collier AY, McMahan K, Yu J, Tostanoski LH, Aguayo R, Ansel J, Chandrashekar A, Patel S, Apraku Bondzie E, Sellers D, Barrett J, Sanborn O, Wan H, Chang A, Anioke T, Nkolola J, Bradshaw C, Jacob-Dolan C, Feldman J, Gebre M, Borducchi EN, Liu J, Schmidt AG, Suscovich T, Linde C, Alter G, Hacker MR, Barouch DH. JAMA. 2021 May 13.
[2] Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) vaccination in pregnancy: Measures of immunity and placental histopathology. Shanes ED, Otero S, Mithal LB, Mupanomunda CA, Miller ES, Goldstein JA. Obstet Gynecol. 2021 May 11.
[3] COVID-19 vaccines while pregnant or breastfeeding. Centers for Disease Control and Prevention.
Links:
COVID-19 Research (NIH)
Barouch Laboratory (Beth Israel Deaconess Medical Center and Harvard Medical School, Boston)
Jeffery Goldstein (Northwestern University Feinberg School of Medicine, Chicago)
NIH Support: National Institute of Allergy and Infectious Diseases; National Cancer Institute, National Institute of Child Health and Human Development; National Center for Advancing Translational Sciences; National Institute of Biomedical Imaging and Bioengineering
Whole-Genome Sequencing Plus AI Yields Same-Day Genetic Diagnoses
Posted on by Dr. Francis Collins

Back in April 2003, when the international Human Genome Project successfully completed the first reference sequence of the human DNA blueprint, we were thrilled to have achieved that feat in just 13 years. Sure, the U.S. contribution to that first human reference sequence cost an estimated $400 million, but we knew (or at least we hoped) that the costs would come down quickly, and the speed would accelerate. How far we’ve come since then! A new study shows that whole genome sequencing—combined with artificial intelligence (AI)—can now be used to diagnose genetic diseases in seriously ill babies in less than 24 hours.
Take a moment to absorb this. I would submit that there is no other technology in the history of planet Earth that has experienced this degree of progress in speed and affordability. And, at the same time, DNA sequence technology has achieved spectacularly high levels of accuracy. The time-honored adage that you can only get two out of three for “faster, better, and cheaper” has been broken—all three have been dramatically enhanced by the advances of the last 16 years.
Rapid diagnosis is critical for infants born with mysterious conditions because it enables them to receive potentially life-saving interventions as soon as possible after birth. In a study in Science Translational Medicine, NIH-funded researchers describe development of a highly automated, genome-sequencing pipeline that’s capable of routinely delivering a diagnosis to anxious parents and health-care professionals dramatically earlier than typically has been possible [1].
While the cost of rapid DNA sequencing continues to fall, challenges remain in utilizing this valuable tool to make quick diagnostic decisions. In most clinical settings, the wait for whole-genome sequencing results still runs more than two weeks. Attempts to obtain faster results also have been labor intensive, requiring dedicated teams of experts to sift through the data, one sample at a time.
In the new study, a research team led by Stephen Kingsmore, Rady Children’s Institute for Genomic Medicine, San Diego, CA, describes a streamlined approach that accelerates every step in the process, making it possible to obtain whole-genome test results in a median time of about 20 hours and with much less manual labor. They propose that the system could deliver answers for 30 patients per week using a single genome sequencing instrument.
Here’s how it works: Instead of manually preparing blood samples, his team used special microbeads to isolate DNA much more rapidly with very little labor. The approach reduced the time for sample preparation from 10 hours to less than three. Then, using a state-of-the-art DNA sequencer, they sequence those samples to obtain good quality whole genome data in just 15.5 hours.
The next potentially time-consuming challenge is making sense of all that data. To speed up the analysis, Kingsmore’s team took advantage of a machine-learning system called MOON. The automated platform sifts through all the data using artificial intelligence to search for potentially disease-causing variants.
The researchers paired MOON with a clinical language processing system, which allowed them to extract relevant information from the child’s electronic health records within seconds. Teaming that patient-specific information with data on more than 13,000 known genetic diseases in the scientific literature, the machine-learning system could pick out a likely disease-causing mutation out of 4.5 million potential variants in an impressive 5 minutes or less!
To put the system to the test, the researchers first evaluated its ability to reach a correct diagnosis in a sample of 101 children with 105 previously diagnosed genetic diseases. In nearly every case, the automated diagnosis matched the opinions reached previously via the more lengthy and laborious manual interpretation of experts.
Next, the researchers tested the automated system in assisting diagnosis of seven seriously ill infants in the intensive care unit, and three previously diagnosed infants. They showed that their automated system could reach a diagnosis in less than 20 hours. That’s compared to the fastest manual approach, which typically took about 48 hours. The automated system also required about 90 percent less manpower.
The system nailed a rapid diagnosis for 3 of 7 infants without returning any false-positive results. Those diagnoses were made with an average time savings of more than 22 hours. In each case, the early diagnosis immediately influenced the treatment those children received. That’s key given that, for young children suffering from serious and unexplained symptoms such as seizures, metabolic abnormalities, or immunodeficiencies, time is of the essence.
Of course, artificial intelligence may never replace doctors and other healthcare providers. Kingsmore notes that 106 years after the invention of the autopilot, two pilots are still required to fly a commercial aircraft. Likewise, health care decisions based on genome interpretation also will continue to require the expertise of skilled physicians.
Still, such a rapid automated system will prove incredibly useful. For instance, this system can provide immediate provisional diagnosis, allowing the experts to focus their attention on more difficult unsolved cases or other needs. It may also prove useful in re-evaluating the evidence in the many cases in which manual interpretation by experts fails to provide an answer.
The automated system may also be useful for periodically reanalyzing data in the many cases that remain unsolved. Keeping up with such reanalysis is a particular challenge considering that researchers continue to discover hundreds of disease-associated genes and thousands of variants each and every year. The hope is that in the years ahead, the combination of whole genome sequencing, artificial intelligence, and expert care will make all the difference in the lives of many more seriously ill babies and their families.
Reference:
[1] Diagnosis of genetic diseases in seriously ill children by rapid whole-genome sequencing and automated phenotyping and interpretation. Clark MM, Hildreth A, Batalov S, Ding Y, Chowdhury S, Watkins K, Ellsworth K, Camp B, Kint CI, Yacoubian C, Farnaes L, Bainbridge MN, Beebe C, Braun JJA, Bray M, Carroll J, Cakici JA, Caylor SA, Clarke C, Creed MP, Friedman J, Frith A, Gain R, Gaughran M, George S, Gilmer S, Gleeson J, Gore J, Grunenwald H, Hovey RL, Janes ML, Lin K, McDonagh PD, McBride K, Mulrooney P, Nahas S, Oh D, Oriol A, Puckett L, Rady Z, Reese MG, Ryu J, Salz L, Sanford E, Stewart L, Sweeney N, Tokita M, Van Der Kraan L, White S, Wigby K, Williams B, Wong T, Wright MS, Yamada C, Schols P, Reynders J, Hall K, Dimmock D, Veeraraghavan N, Defay T, Kingsmore SF. Sci Transl Med. 2019 Apr 24;11(489).
Links:
DNA Sequencing Fact Sheet (National Human Genome Research Institute/NIH)
Genomics and Medicine (NHGRI/NIH)
Genetic and Rare Disease Information Center (National Center for Advancing Translational Sciences/NIH)
Stephen Kingsmore (Rady Children’s Institute for Genomic Medicine, San Diego, CA)
NIH Support: National Institute of Child Health and Human Development; National Human Genome Research Institute; National Center for Advancing Translational Sciences
Preeclampsia: Study Highlights Need for More Effective Treatment, Prevention
Posted on by Dr. Francis Collins
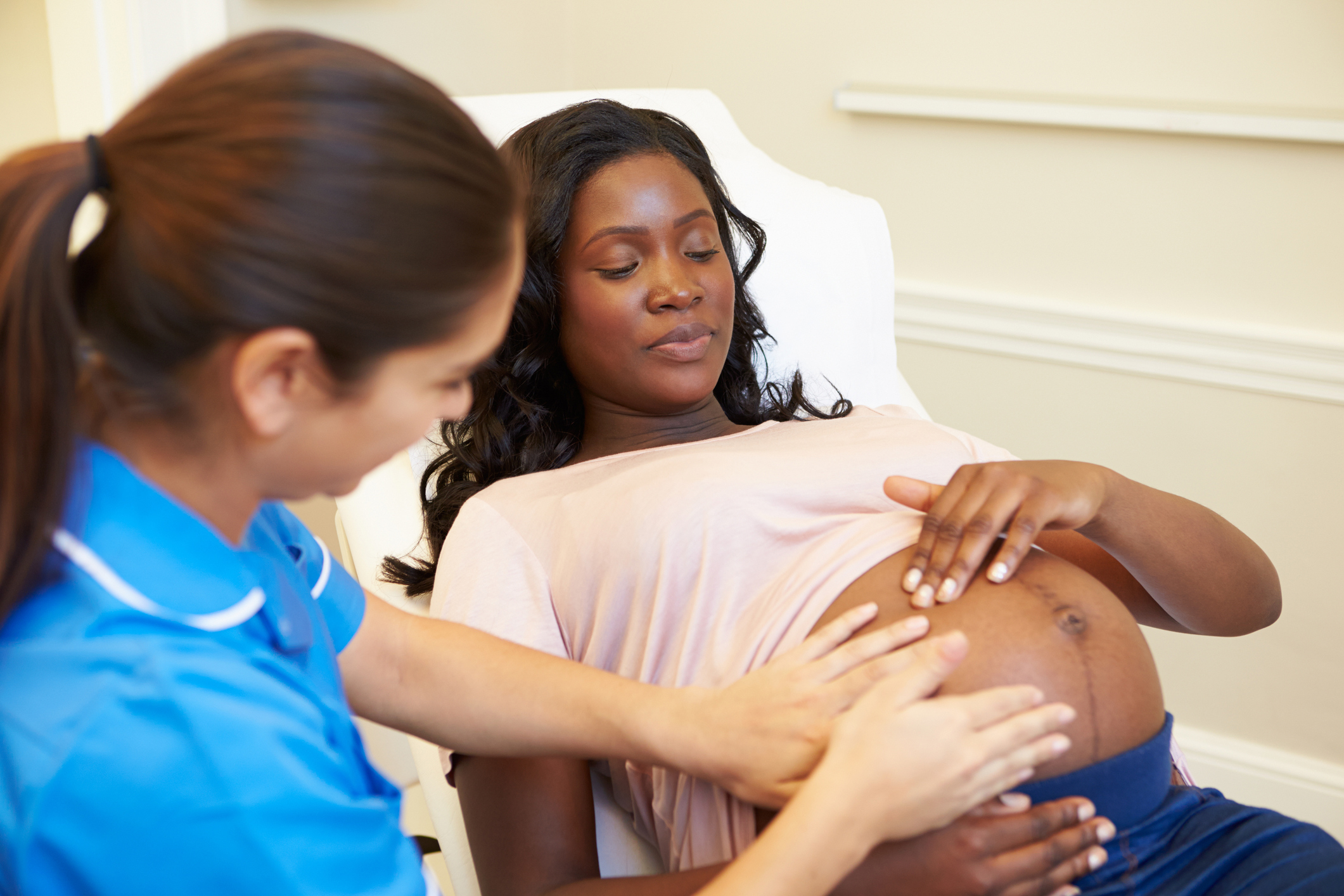

Thinkstock
It’s well known that preeclampsia, a condition characterized by a progressive rise in a pregnant woman’s blood pressure and appearance of protein in the urine, can have negative, even life-threatening impacts on the health of both mother and baby. Now, NIH-funded researchers have documented that preeclampsia is also taking a very high toll on our nation’s economic well-being. In fact, their calculations show that, in 2012 alone, preeclampsia-related care cost the U.S. health care system more than $2 billion.
These findings are especially noteworthy because preeclampsia rates in the United States have been steadily rising over the past 30 years, fueled in part by increases in average maternal age and weight. This highlights the urgent need for more research to develop new and more effective strategies to protect the health of all mothers and their babies.
Autism Spectrum Disorder: Progress Toward Earlier Diagnosis
Posted on by Dr. Francis Collins

Stockbyte
Research shows that the roots of autism spectrum disorder (ASD) generally start early—most likely in the womb. That’s one more reason, on top of a large number of epidemiological studies, why current claims about the role of vaccines in causing autism can’t be right. But how early is ASD detectable? It’s a critical question, since early intervention has been shown to help limit the effects of autism. The problem is there’s currently no reliable way to detect ASD until around 18–24 months, when the social deficits and repetitive behaviors associated with the condition begin to appear.
Several months ago, an NIH-funded team offered promising evidence that it may be possible to detect ASD in high-risk 1-year-olds by shifting attention from how kids act to how their brains have grown [1]. Now, new evidence from that same team suggests that neurological signs of ASD might be detectable even earlier.
Peanut Allergy: Early Exposure Is Key to Prevention
Posted on by Dr. Francis Collins
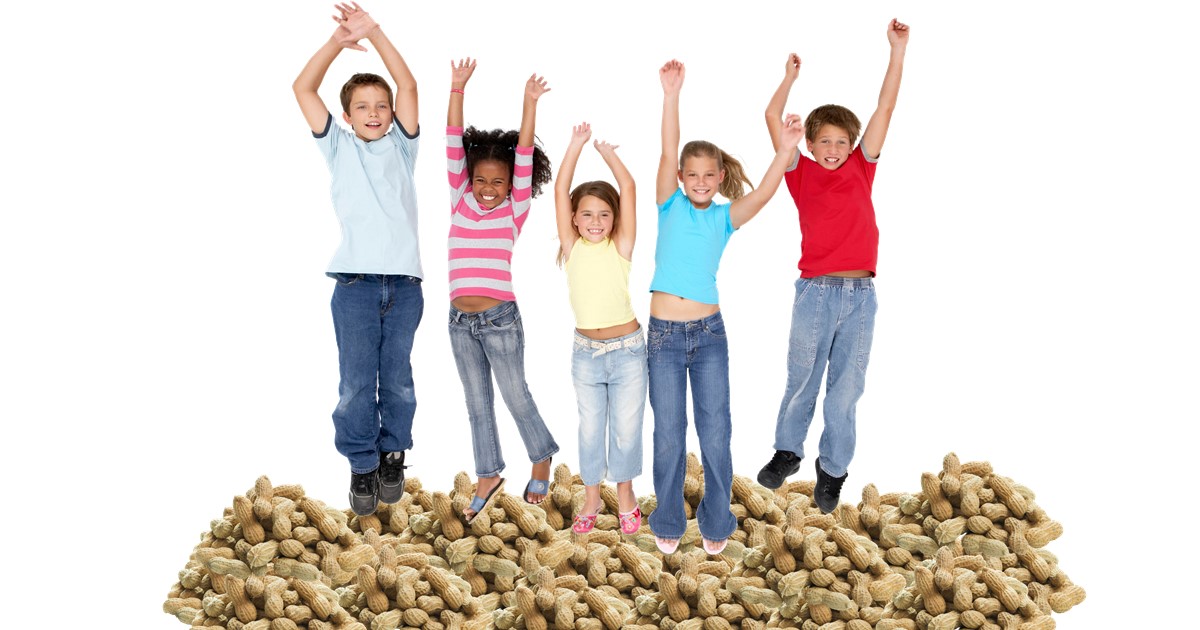

Credit: Thinkstock (BananaStock, Kenishirotie)
With peanut allergy on the rise in the United States, you’ve probably heard parents strategizing about ways to keep their kids from developing this potentially dangerous condition. But is it actually possible to prevent peanut allergy, and, if so, how do you go about doing it?
There’s an entirely new strategy emerging now! A group representing 26 professional organizations, advocacy groups, and federal agencies, including the National Institutes of Health (NIH), has just issued new clinical guidelines aimed at preventing peanut allergy [1]. The guidelines suggest that parents should introduce most babies to peanut-containing foods around the time they begin eating other solid foods, typically 4 to 6 months of age. While early introduction is especially important for kids at particular risk for developing allergies, it is also recommended that high-risk infants—those with a history of severe eczema and/or egg allergy—undergo a blood or skin-prick test before being given foods containing peanuts. The test results can help to determine how, or even if, peanuts should be introduced in the youngsters’ diets.
Largest Study Yet Shows Mother’s Smoking Changes Baby’s Epigenome
Posted on by Dr. Francis Collins

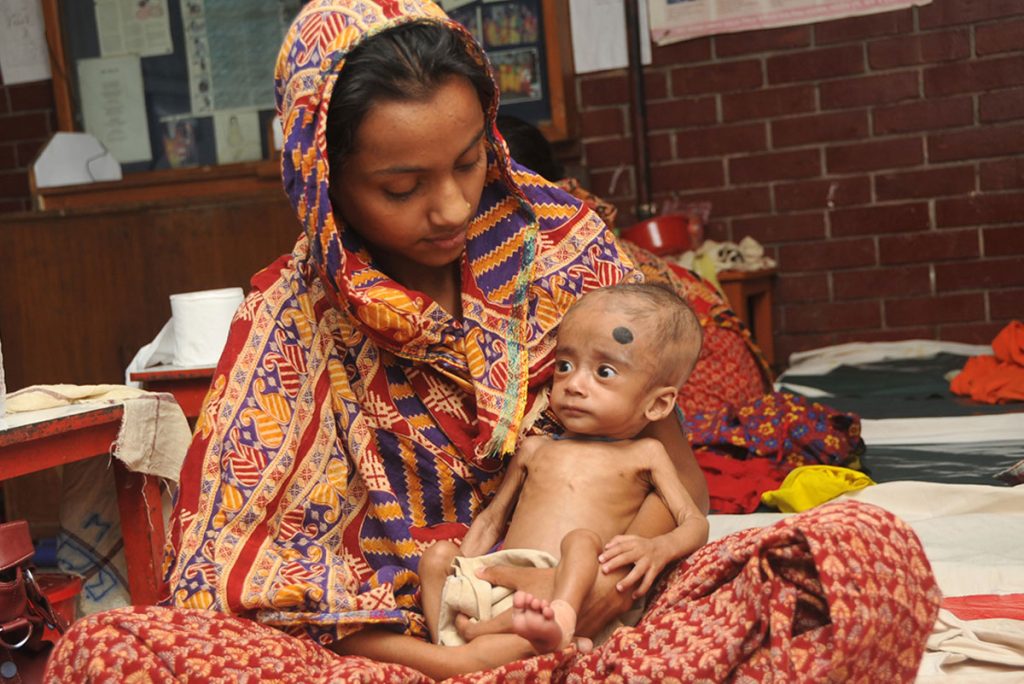
Credit: Daniel Berehulak/Getty Images
Despite years of public health campaigns warning of the dangers of smoking when pregnant, many women are unaware of the risk or find themselves unable to quit. As a result, far too many babies are still being exposed in the womb to toxins that enter their mothers’ bloodstreams when they inhale cigarette smoke. Among the many infant and child health problems that have been linked to maternal smoking are premature birth, low birth weight, asthma, reduced lung function, sudden infant death syndrome (SIDS), and cleft lip and/or palate.
Now, a large international study involving NIH-supported researchers provides a biological mechanism that may explain how exposure to cigarette toxins during fetal development can produce these health problems [1]. That evidence centers on the impact of the toxins on the epigenome of the infant’s body tissues. The epigenome refers to chemical modifications of DNA (particularly methylation of cytosines), as well as proteins that bind to DNA and affect its function. The genome of an individual is the same in all cells of their body, but the epigenome determines whether genes are turned on or off in particular cells. The study found significant differences between the epigenetic patterns of babies born to women who smoked during pregnancy and those born to non-smokers, with many of the differences affecting genes known to play key roles in the development of the lungs, face, and nervous system.
Creative Minds: A Baby’s Eye View of Language Development
Posted on by Dr. Francis Collins
 If you are a fan of wildlife shows, you’ve probably seen those tiny video cameras rigged to animals in the wild that provide a sneak peek into their secret domains. But not all research cams are mounted on creatures with fur, feathers, or fins. One of NIH’s 2014 Early Independence Award winners has developed a baby-friendly, head-mounted camera system (shown above) that captures the world from an infant’s perspective and explores one of our most human, but still imperfectly understood, traits: language.
If you are a fan of wildlife shows, you’ve probably seen those tiny video cameras rigged to animals in the wild that provide a sneak peek into their secret domains. But not all research cams are mounted on creatures with fur, feathers, or fins. One of NIH’s 2014 Early Independence Award winners has developed a baby-friendly, head-mounted camera system (shown above) that captures the world from an infant’s perspective and explores one of our most human, but still imperfectly understood, traits: language.
Elika Bergelson, a young researcher at the University of Rochester in New York, wants to know exactly how and when infants acquire the ability to understand spoken words. Using innovative camera gear and other investigative tools, she hopes to refine current thinking about the natural timeline for language acquisition. Bergelson also hopes her work will pay off in a firmer theoretical foundation to help clinicians assess children with poor verbal skills or with neurodevelopmental conditions that impair information processing, such as autism spectrum disorders.


 For centuries, people have yearned for an elixir capable of restoring youth to their aging bodies and minds. It sounds like pure fantasy, but, in recent years, researchers have shown that the blood of young mice can exert a regenerative effect
For centuries, people have yearned for an elixir capable of restoring youth to their aging bodies and minds. It sounds like pure fantasy, but, in recent years, researchers have shown that the blood of young mice can exert a regenerative effect 
