skin
Can Bioprinted Skin Substitutes Replace Traditional Grafts for Treating Burn Injuries and Other Serious Skin Wounds?
Posted on by Lawrence Tabak, D.D.S., Ph.D.
Each year in the U.S., more than 500,000 people receive treatment for burn injuries and other serious skin wounds.1 To close the most severe wounds with less scarring, doctors often must surgically remove skin from one part of a person’s body and use it to patch the injured site. However, this is an intensive process, and some burn patients with extensive skin loss do not have sufficient skin available for grafting. Scientists have been exploring ways to repair these serious skin wounds without skin graft surgery.
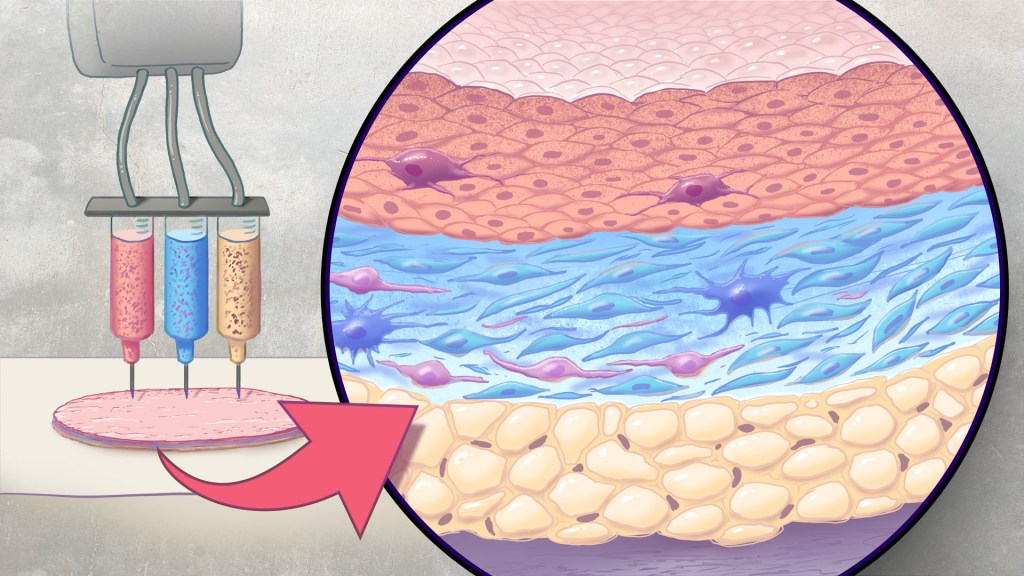
An NIH-funded team recently showed that bioprinted skin substitutes may serve as a promising alternative to traditional skin grafts in preclinical studies reported in Science Translational Medicine.2 The approach involves a portable skin bioprinter system that deposits multiple layers of skin directly into a wound. The recent findings add to evidence that bioprinting technology can successfully regenerate human-like skin to allow healing. While this approach has yet to be tested in people, it confirms that such technologies already can produce skin constructs with the complex structures and multiple cell types present in healthy human skin.
This latest work comes from a team led by Adam Jorgensen and Anthony Atala at Wake Forest School of Medicine’s Wake Forest Institute for Regenerative Medicine, Winston-Salem, NC. Members of the Atala lab and their colleagues had earlier shown it was possible to isolate two major skin cell types found in the skin’s outer (epidermis) and middle (dermis) layers from a small biopsy of healthy skin, expand the number of cells in the lab and then deliver the cells directly into an injury using a specially designed bioprinter.3 Using integrated imaging technology to scan a wound, computer software “prints” cells right into an injury, mimicking two of our skin’s three natural layers.
In the new study, Atala’s team has gone even further to construct skin substitutes that mimic the structure of human skin and that include six primary human skin cell types. They then used their bioprinter to produce skin constructs with all three layers found in healthy human skin: epidermis, dermis, and hypodermis.
To put their skin substitutes to the test, they first transplanted them into mice. Their studies showed that the bioprinted skin encouraged the rapid growth of new blood vessels and had other features of normal-looking, healthy skin. The researchers were able to confirm that their bioprinted skin implants successfully integrated into the animals’ regenerated skin to speed healing.
Studies in a pig model of wound healing added to evidence that such bioprinted implants can successfully repair full-thickness wounds, meaning those that extend through all three layers of skin. The bioprinted skin patches allowed for improved wound healing with less scarring. They also found that the bioprinted grafts encouraged activity in the skin from genes known to play important roles in wound healing.
It’s not yet clear if this approach will work as well in the clinic as it does in the lab. To make it feasible, the researchers note there’s a need for improved approaches to isolating and expanding the needed skin cell types. Nevertheless, these advances come as encouraging evidence that bioprinted skin substitutes could one day offer a promising alternative to traditional skin grafts with the capacity to help even more people with severe burns or other wounds.
References:
[1] Burn Incidence Fact Sheet. American Burn Association
[2] AM Jorgensen, et al. Multicellular bioprinted skin facilitates human-like skin architecture in vivo. Science Translational Medicine DOI: 10.1126/scitranslmed.adf7547 (2023).
[3] M Albanna, et al. In Situ Bioprinting of Autologous Skin Cells Accelerates Wound Healing of Extensive Excisional Full-Thickness Wounds. Scientific Reports DOI: 10.1038/s41598-018-38366-w (2019).
NIH Support: National Institute of Arthritis and Musculoskeletal and Skin Diseases
Mapping Immune Cell “Neighborhoods” in Psoriasis to Understand its Course
Posted on by Lawrence Tabak, D.D.S., Ph.D.

“Location, location, location.” While most of us know this phrase as a real estate adage, location—specifically that of various cell types—is becoming a key area of investigation in studying human disease. New techniques are enabling scientists to understand where certain cells are with respect to one another and how changes in their activity may affect your overall health.
In one recent example of the power of this approach, NIH-funded researchers [1] used a sophisticated method to map immune cells within human skin to get a more detailed picture of psoriasis, a common, chronic disease in which the immune system becomes overactive leading to skin inflammation. People with psoriasis develop patches of itchy, red, and flaky lesions on their skin, which can be mild to severe. For reasons that aren’t entirely clear, they’re also at higher risk for developing a wide range of other health conditions, including a unique form of arthritis known as psoriatic arthritis, diabetes, mental health issues, heart problems, and more.
The hope is that these newly drawn, precise maps of cellular “neighborhoods” in human skin will help chart the precise course of this disease to understand better the differences between mild and more severe forms. They may also yield important clues as to why people with psoriasis develop other health problems more often than people without psoriasis.
In the new study, a team including Jose Scher and Shruti Naik, NYU Langone, New York, analyzed immune cells within 25 skin samples from 14 volunteers, including those with active psoriasis, those with psoriasis but no active lesions, and people with healthy skin who do not have psoriasis. The researchers relied on a sophisticated approach called spatial transcriptomics [2] to map out what happens at the single-cell level within the samples.
In earlier approaches to single-cell analysis, researchers first would separate cells from the tissue they came from. While they could measure gene activity within those cells at the individual level, they couldn’t put things back together to see how they all fit. With spatial transcriptomics, it’s now possible to molecularly profile single cells to measure their activity in a tissue sample while also mapping their locations with respect to other cells.
The new study led to some intriguing findings. For instance, certain immune cells, specifically B cells, moved to the upper layers of the skin during active disease. That’s notable because prior studies had been unable to capture B cells in the skin adequately, and these cells are thought to play an important role in the disease.
Interestingly, the spatial cellular maps revealed inflammatory regions in both actively inflamed skin and in skin that appeared healthy. This finding highlights the fact that the inflammation that goes with psoriasis can affect the skin, and likely other parts of the body, in ways that aren’t easily observed. In future studies, the researchers want to explore how the presence of psoriasis and its underlying changes in immune cell activity may influence other organs and tissues beneath the skin.
Their fine-scale maps also showed increased gene activity in dozens of molecular pathways that are tied to metabolism and the control of lipid levels. That’s especially interesting because these factors are known to go awry in diabetes and heart conditions, which happen more often in people with psoriasis compared to those without. They also could see in their maps that this altered activity sometimes occurred in clear skin distant from any apparent lesions.
Having discovered such signals with potential consequences for other parts of the body, the researchers report that they’re working to understand how inflammatory immune cells and processes in the skin may lead to more widespread disease processes that affect other parts of the body. They plan to conduct similar studies in larger groups of people with and without active psoriasis lesions and studies following individuals with psoriasis over time. They’ll also explore questions about why people respond differently to the same anti-inflammatory treatment regimens.
To speed the process of discovery, they’ve made their maps and associated data freely available as a resource for the scientific community. About 7.5 million adults in the U.S. and millions more worldwide have psoriasis and associated psoriatic conditions [3]. The hope is that these maps will one day help to steer them toward a healthier future.
References:
[1] Spatial transcriptomics stratifies psoriatic disease severity by emergent cellular ecosystems. Castillo RL, Sidhu I, Dolgalev I, Chu T, Prystupa A, Subudhi I, Yan D, Konieczny P, Hsieh B, Haberman RH, Selvaraj S, Shiomi T, Medina R, Girija PV, Heguy A, Loomis CA, Chiriboga L, Ritchlin C, Garcia-Hernandez ML, Carucci J, Meehan SA, Neimann AL, Gudjonsson JE, Scher JU, Naik S. Sci Immunol. 2023 Jun 8;8(84):eabq7991. doi: 10.1126/sciimmunol.abq7991.
[2] Method of the Year: spatially resolved transcriptomics. Marx V. Nat Methods. 2021 Jan;18(1):9-14. doi: 10.1038/s41592-020-01033-y.
[3] Psoriasis Prevalence in Adults in the United States. Armstrong AW, Mehta MD, Schupp CW, Gondo GC, Bell SJ, Griffiths CEM. JAMA Dermatol. 2021 Aug 1;157(8):940-946. doi: 10.1001/jamadermatol.2021.2007.
Links:
Psoriasis (National Institute of Arthritis and Musculoskeletal and Skin Diseases/NIH)
Jose Scher (NYU Langone Health, New York, NY)
Shruti Naik (NYU Langone Health, New York, NY)
NIH Support: National Cancer Institute, National Institute of Arthritis and Musculoskeletal and Skin Diseases, National Center for Advancing Translational Sciences, National Institute of Allergy and Infectious Diseases
A More Precise Way to Knock Out Skin Rashes
Posted on by Lawrence Tabak, D.D.S., Ph.D.
The NIH is committed to building a new era in medicine in which the delivery of health care is tailored specifically to the individual person, not the hypothetical average patient as is now often the case. This new era of “precision medicine” will transform care for many life-threatening diseases, including cancer and chronic kidney disease. But what about non-life-threatening conditions, like the aggravating rash on your skin that just won’t go away?
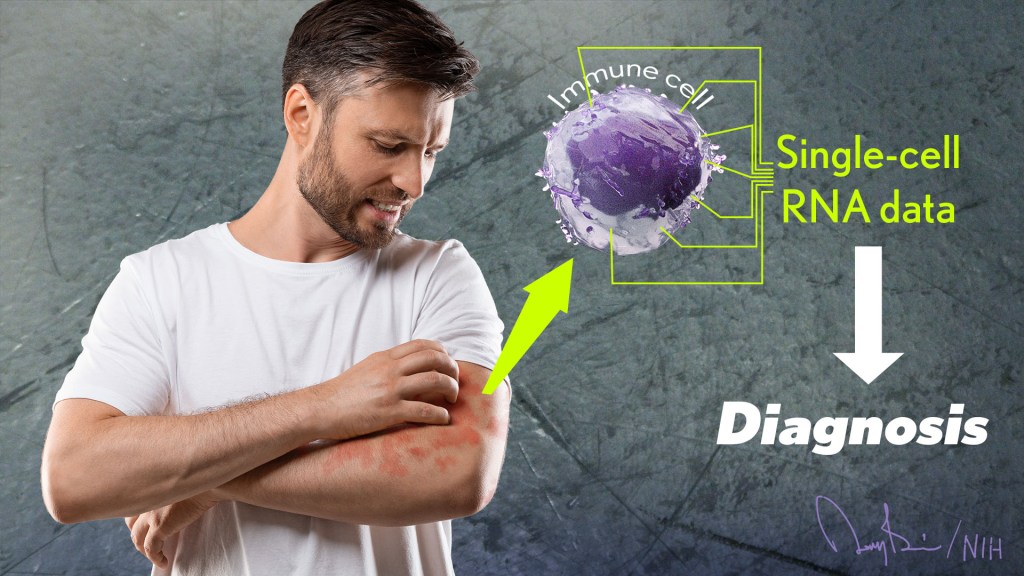
Recently, researchers published a proof-of-principle paper in the journal Science Immunology demonstrating just how precision medicine for inflammatory skin rashes might work [1]. While more research is needed to build out and further refine the approach, the researchers show it’s now technologically possible to extract immune cells from a patient’s rash, read each cell’s exact inflammatory features, and relatively quickly match them online to the right anti-inflammatory treatment to stop the rash.
The work comes from a NIH-funded team led by Jeffrey Cheng and Raymond Cho, University of California, San Francisco. The researchers focused their attention on two inflammatory skin conditions: atopic dermatitis, the most common type of eczema, which flares up periodically to make skin red and itchy, and psoriasis vulgaris. Psoriasis causes skin cells to build up and form a scaly rash and dry, itchy patches. Together, atopic dermatitis and psoriasis vulgaris affect about 10 percent of U.S. adults.
While the rashes caused by the two conditions can sometimes look similar, they are driven by different sets of immune cells and underlying inflammatory responses. For that reason, distinct biologic therapies, based on antibodies and proteins made from living cells, are now available to target and modify the specific immune pathways underlying each condition.
While biologic therapies represent a major treatment advance for these and other inflammatory conditions, they can miss their targets. Indeed, up to half of patients don’t improve substantially on biologics. Part of the reason for that lack of improvement is because doctors don’t have the tools they need to make firm diagnoses based on what precisely is going on in the skin at the molecular and cellular levels.
To learn more in the new study, the researchers isolated immune cells, focusing primarily on T cells, from the skin of 31 volunteers. They then sequenced the RNA of each cell to provide a telltale portrait of its genomic features. This “single-cell analysis” allowed them to capture high-resolution portraits of 41 different immune cell types found in individual skin samples. That’s important because it offers a much more detailed understanding of changes in the behavior of various immune cells that might have been missed in studies focused on larger groupings of skin cells, representing mixtures of various cell types.
Of the 31 volunteers, seven had atopic dermatitis and eight had psoriasis vulgaris. Three others were diagnosed with other skin conditions, while six had an indeterminate rash with features of both atopic dermatitis and psoriasis vulgaris. Seven others were healthy controls.
The team produced molecular signatures of the immune cells. The researchers then compared the signatures from the hard-to-diagnose rashes to those of confirmed cases of atopic dermatitis and psoriasis. They wanted to see if the signatures could help to reach clearer diagnoses.
The signatures revealed common immunological features as well as underlying differences. Importantly, the researchers found that the signatures allowed them to move forward and classify the indeterminate rashes. The rashes also responded to biologic therapies corresponding to the individuals’ new diagnoses.
Already, the work has identified molecules that help to define major classes of human inflammatory skin diseases. The team has also developed computer tools to help classify rashes in many other cases where the diagnosis is otherwise uncertain.
In fact, the researchers have launched a pioneering website called RashX. It is enabling practicing dermatologists and researchers around the world to submit their single-cell RNA data from their difficult cases. Such analyses are now being done at a small, but growing, number of academic medical centers.
While precision medicine for skin rashes has a long way to go yet before reaching most clinics, the UCSF team is working diligently to ensure its arrival as soon as scientifically possible. Indeed, their new data represent the beginnings of an openly available inflammatory skin disease resource. They ultimately hope to generate a standardized framework to link molecular features to disease prognosis and drug response based on data collected from clinical centers worldwide. It’s a major effort, but one that promises to improve the diagnosis and treatment of many more unusual and long-lasting rashes, both now and into the future.
Reference:
[1] Classification of human chronic inflammatory skin disease based on single-cell immune profiling. Liu Y, Wang H, Taylor M, Cook C, Martínez-Berdeja A, North JP, Harirchian P, Hailer AA, Zhao Z, Ghadially R, Ricardo-Gonzalez RR, Grekin RC, Mauro TM, Kim E, Choi J, Purdom E, Cho RJ, Cheng JB. Sci Immunol. 2022 Apr 15;7(70):eabl9165. {Epub ahead of publication]
Links:
The Promise of Precision Medicine (NIH)
Atopic Dermatitis (National Institute of Arthritis and Musculoskeletal and Skin Diseases /NIH)
Psoriasis (NIAMS/NIH)
RashX (University of California, San Francisco)
Raymond Cho (UCSF)
Jeffrey Cheng (UCSF)
NIH Support: National Institute of Arthritis and Musculoskeletal and Skin Diseases; National Center for Advancing Translational Sciences
Unraveling the Role of the Skin Microbiome in Health and Disease
Posted on by Lindsey A. Criswell, M.D., M.P.H., D.Sc., National Institute of Arthritis and Musculoskeletal and Skin Diseases

Human skin is home to diverse ecosystems including bacteria, viruses, and fungi. These microbial communities comprise hundreds of species and are collectively known as the skin microbiome. The skin microbiome is thought to play a vital role in fending off disease-causing microorganisms (pathogens), boosting barrier protection, and aiding immune defenses.
Maintaining a balanced skin microbiome involves a complex and dynamic interplay among microorganisms, immune cells, skin cells, and other factors. In general, bacteria far outnumber viral, fungal, or other microbial species on the skin. Bacterial communities, which are strongly influenced by conditions such as skin moisture, temperature, and pH, vary widely across the body. For example, facial cheek skin hosts mostly Cutibacterium along with a bit of the skin fungus Malassezia. The heel is colonized by different types of bacteria including Staphylococcus and Corynebacteria.
In some diseases, such as acne and eczema, the skin microbiome is altered. Typically, this means an increase in pathogenic microorganisms and a decrease in beneficial ones. An altered skin microbiome can also be associated with inflammation, severe disease symptoms, and changes in the human immune system.
Heidi H. Kong is working to understand the role of the skin microbiome in health and disease. She is a senior investigator in the Intramural Research Program at NIH’s National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS) and an adjunct investigator at NIH’s National Cancer Institute (NCI).
More than a decade ago, Kong and Julie A. Segre, an intramural researcher at NIH’s National Human Genome Research Institute, analyzed the microbial makeup of healthy individuals. Kong swabbed the skin of these healthy volunteers in 20 different sites, from the forehead to the toenail. The study revealed that the surface of the human body provides various environmental niches, depending on whether the skin is moist, dry, or sebaceous (oily). Different bacterial species predominate in each niche. Kong and Segre were particularly interested in body areas that have predilections for disease. For example, psoriasis is often found on the outside of elbows and knees, and the back of the scalp.
Earlier this year, Kong and Segre published another broad analysis of the human skin microbiome [1] in collaboration with scientists at the European Molecular Biology Laboratory, European Bioinformatics Institute (EMBL-EBI), United Kingdom. This new catalog, called the Skin Microbial Genome Collection, is thought to identify about 85 percent of the microorganisms present on healthy skin from 19 body sites. It documents more than 600 bacterial species—including 174 that were discovered during the study—as well as more than 6,900 viruses and some fungi, including three newly discovered species.
Kong’s work has provided compelling evidence that the human immune system plays a role in shaping the skin microbiome. In 2018, she, Segre, and colleagues from the intramural programs of NCI and NIH’s National Institute of Allergy and Infectious Diseases analyzed skin from eight different sites on 27 people with a rare primary immunodeficiency disease known as DOCK8 deficiency [2].
People with the condition have recurrent infections in the skin, sinuses, and airways, and are susceptible to different cancers. Kong and colleagues found that the skin of people with DOCK8 deficiency contains significantly more DNA viruses (90 percent of the skin microbiome on average) than people without the condition (6 or 7 percent of the skin microbiome).
Other researchers are hoping to leverage features of the microbiome to develop targeted therapies for skin diseases. Richard L. Gallo, a NIAMS grantee at the University of California, San Diego, is currently focused on acne and eczema (also called atopic dermatitis). Acne is associated with certain strains of Cutibacterium acnes (C. acnes, formerly called Propionibacterium acnes or P. acnes). Eczema is often associated with Staphylococcus aureus (S. aureus).
Severe cases of acne and eczema are commonly treated with broad-spectrum antibiotics, which wipe out most of the bacteria, including beneficial species. The goal of microbiome-targeted therapy is to kill only the disease-associated bacteria and avoid increasing the risk that some strains will develop antibiotic resistance.
In 2020, Gallo and colleagues identified a strain of Staphylococcus capitis from healthy human skin (S. capitis E12) that selectively inhibits the growth of C. acnes without negatively impacting other bacteria or human skin cells [3]. S. capitis E12 produces four different toxins that act together to target C. acnes. The research team created an extract of the four toxins and tested it using animal models. In most cases, the extract was more potent at killing C. acnes—including acne-associated strains—than several commonly prescribed antibiotics (erythromycin, tetracycline, and clindamycin). And, unlike antibiotics, the extract does not appear to promote drug-resistance, at least for the 20 generations observed by the researchers.
Eczema is a chronic, relapsing disease characterized by skin that is dry, itchy, inflamed, and prone to infection, including by pathogens such as S. aureus and herpes virus. Although the cause of eczema is unknown, the condition is associated with human genetic mutations, disruption of the skin’s barrier, inflammation-triggering allergens, and imbalances in the skin microbiome.
In 2017, Gallo’s research team discovered that, in healthy human skin, certain strains of Staphylococcus hominis and Staphylococcus epidermis produce potent antimicrobial molecules known as lantibiotics [4]. These beneficial strains are far less common on the skin of people with eczema. The lantibiotics work synergistically with LL-37, an antimicrobial molecule produced by the human immune system, to selectively kill S. aureus, including methicillin-resistant strains (MRSA).
Gallo and his colleagues then examined the safety and therapeutic potential of these beneficial strains isolated from the human skin microbiome. In animal tests, strains of S. hominis and S. epidermis that produce lantibiotics killed S. aureus and blocked production of its toxin.
Gallo’s group has now expanded their work to early studies in humans. In 2021, two independent phase 1 clinical trials [5,6] conducted by Gallo and his colleagues investigated the effects of these strains on people with eczema. These double-blind, placebo-controlled trials involved one-week of topical application of beneficial bacteria to the forearm of adults with S. aureus-positive eczema. The results demonstrated that the treatment was safe, showed a significant decrease in S. aureus, and improved eczema symptoms in most patients. This is encouraging news for those hoping to develop microbiome-targeted therapy for inflammatory skin diseases.
As research on the skin microbiome advances on different fronts, it will provide deeper insight into the multi-faceted microbial communities that are so critical to health and disease. One day, we may even be able to harness the microbiome as a source of therapeutics to alleviate inflammation, promote wound healing, or suppress certain skin cancers.
References:
[1] Integrating cultivation and metagenomics for a multi-kingdom view of skin microbiome diversity and functions. Saheb Kashaf S, Proctor DM, Deming C, Saary P, Hölzer M; NISC Comparative Sequencing Program, Taylor ME, Kong HH, Segre JA, Almeida A, Finn RD. Nat Microbiol. 2022 Jan;7(1):169-179.
[2] Expanded skin virome in DOCK8-deficient patients. Tirosh O, Conlan S, Deming C, Lee-Lin SQ, Huang X; NISC Comparative Sequencing Program, Su HC, Freeman AF, Segre JA, Kong HH. Nat Med. 2018 Dec;24(12):1815-1821.
[3] Identification of a human skin commensal bacterium that selectively kills Cutibacterium acnes. O’Neill AM, Nakatsuji T, Hayachi A, Williams MR, Mills RH, Gonzalez DJ, Gallo RL. J Invest Dermatol. 2020 Aug;140(8):1619-1628.e2.
[4] Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Nakatsuji T, Chen TH, Narala S, Chun KA, Two AM, Yun T, Shafiq F, Kotol PF, Bouslimani A, Melnik AV, Latif H, Kim JN, Lockhart A, Artis K, David G, Taylor P, Streib J, Dorrestein PC, Grier A, Gill SR, Zengler K, Hata TR, Leung DY, Gallo RL. Sci Transl Med. 2017 Feb 22;9(378):eaah4680.
[5] Development of a human skin commensal microbe for bacteriotherapy of atopic dermatitis and use in a phase 1 randomized clinical trial. Nakatsuji T, Hata TR, Tong Y, Cheng JY, Shafiq F, Butcher AM, Salem SS, Brinton SL, Rudman Spergel AK, Johnson K, Jepson B, Calatroni A, David G, Ramirez-Gama M, Taylor P, Leung DYM, Gallo RL. Nat Med. 2021 Apr;27(4):700-709.
[6] Use of autologous bacteriotherapy to treat Staphylococcus aureus in patients with atopic dermatitis: A randomized double-blind clinical trial. Nakatsuji T, Gallo RL, Shafiq F, Tong Y, Chun K, Butcher AM, Cheng JY, Hata TR. JAMA Dermatol. 2021 Jun 16;157(8):978-82.
Links:
Acne (National Institute of Arthritis and Musculoskeletal and Skin Diseases/NIH)
Atopic Dermatitis (NIAMS)
Cutaneous Microbiome and Inflammation Laboratory, Heidi Kong (NIAMS)
Julie Segre (National Human Genome Research Institute/NIH)
Gallo Lab (University of California, San Diego)
[Note: Acting NIH Director Lawrence Tabak has asked the heads of NIH’s Institutes and Centers (ICs) to contribute occasional guest posts to the blog to highlight some of the cool science that they support and conduct. This is the fifth in the series of NIH IC guest posts that will run until a new permanent NIH director is in place.]
Capturing the Extracellular Matrix in 3D Color
Posted on by Dr. Francis Collins
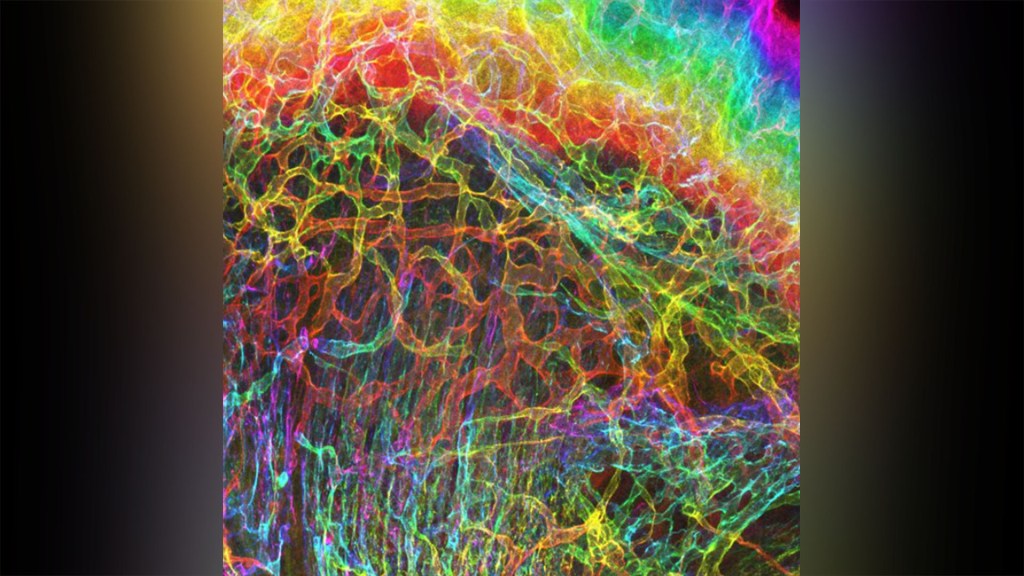
For experienced and aspiring shutterbugs alike, sometimes the best photo in the bunch turns out to be a practice shot. That’s also occasionally true in the lab when imaging cells and tissues, and it’s the story behind this spectacular image showing the interface of skin and muscle during mammalian development.
Here you see an area of the mouse forelimb located near a bone called the humerus. This particular sample was labeled for laminin, a protein found in the extracellular matrix (ECM) that undergirds cells and tissues to give them mechanical and biochemical support. Computer algorithms were used to convert the original 2D confocal scan into a 3D image, and colorization was added to bring the different layers of tissue into sharper relief.
Skin tissue (bright red and yellow) is located near the top of the image; blood vessels (paler red, orange, and yellow) are in the middle and branching downward; and muscle (green, blue, and purple) makes up the bottom layer.
The image was created by Sarah Lipp, a graduate student in the NIH-supported tissue engineering lab of Sarah Calve. The team focuses on tissue interfaces to better understand the ECM and help devise strategies to engineer musculoskeletal tissues, such as tendon and cartilage.
In February 2020, Lipp was playing around with some new software tools for tissue imaging. Before zeroing in on her main target—the mouse’s myotendinous junction, where muscle transfers its force to tendon, Lipp snapped this practice shot of skin meeting muscle. After processing the practice shot with a color-projecting macro in an image processing tool called Fiji, she immediately liked what she saw.
So, Lipp tweaked the color a bit more and entered the image in the 2020 BioArt Scientific Image & Video Competition, sponsored by the Federation of American Societies for Experimental Biology, Bethesda, MD. Last December, the grad student received the good news that her practice shot had snagged one of the prestigious contest’s top awards.
But she’s not stopping there. Lipp is continuing to pursue her research interests at the University of Colorado, Boulder, where the Calve lab recently moved from Purdue University, West Lafayette, IN. Here’s wishing her a career filled with more great images—and great science!
Links:
Muscle and Bone Diseases (National Institute of Arthritis and Musculoskeletal and Skin Diseases/NIH)
Musculoskeletal Extracellular Matrix Laboratory (University of Colorado, Boulder)
BioArt Scientific Image & Video Competition (Federation of American Societies for Experimental Biology, Bethesda, MD)
NIH Support: National Institute of Arthritis and Musculoskeletal and Skin Diseases
How to Heal Skin Without the Scars
Posted on by Dr. Francis Collins

Most of us can point to a few unwanted scars on our bodies. Every scar tells a story, but people are spending billions of dollars each year trying to hide or get rid of them [1]. What if there was a way to get the wounds on our skin to heal without scarring in the first place?
In a recent paper in the journal Science, a team of NIH-supported researchers has taken an important step in this direction. Working with mice, the researchers deciphered some of the key chemical and physical signals that cause certain skin cells to form tough, fibrous scars while healing a wound [2]. They also discovered how to reprogram them with a topical treatment and respond to injuries more like fetal skin cells, which can patch up wounds in full, regrowing hair, glands, and accessory structures of the skin, and all without leaving a mark.
Of course, mice are not humans. Follow-up research is underway to replicate these findings in larger mammals with skin that’s tighter and more akin to ours. But if the preclinical data hold up, the researchers say they can test in future human clinical trials the anti-scarring drug used in the latest study, which has been commercially available for two decades to treat blood vessel disorders in the eye.
The work comes from Michael Longaker, Shamik Mascharak, and colleagues, Stanford Medicine, Palo Alto, CA. But, to be more precise, the work began with a research project that Longaker was given back in 1987, while a post doc in the lab of Michael Harrison, University of California, San Francisco.
Harrison, a surgeon then performing groundbreaking prenatal surgery, noticed that babies born after undergoing surgery in the womb healed from their surgeries without any scarring. He asked his postdoc to find out why, and Longaker has been trying to answer that question and understand scar formation ever since.
Longaker and his Stanford colleague Geoffrey Gurtner suspected that the difference between healing inside and outside the womb had something to do with tension. Inside the womb, the skin of the unborn is bathed in fluid and develops in a soft, tension-free state. Outside the womb, human skin is exposed to continuous environmental stresses and must continuously remodel and grow to remain viable, which creates a high level of skin tension.
Following up on Longaker and Gurtner’s suspicion, Mascharak found in a series of mouse experiments that a particular class of fibroblast, a type of cell in skin and other connective tissues, activates a gene called Engrailed-1 during scar formation [3]. To see if mechanical stress played a role in this process, Mascharak and team grew mouse fibroblast cells on either a soft, stress-free gel or on a stiff plastic dish that produced mechanical strain. Importantly, they also tried growing the fibroblasts on the same strain-inducing plastic, but in the presence of a chemical that blocked the mechanical-strain signal.
Their studies showed that fibroblasts grown on the tension-free gel didn’t activate the scar-associated genetic program, unlike fibroblasts growing on the stress-inducing plastic. With the chemical that blocked the cells’ ability to sense the mechanical strain, Engrailed-1 didn’t get switched on either.
They also showed the opposite. When tension was applied to healing surgical incisions in mice, it led to an increase in the number of those fibroblast cells expressing Engrailed-1 and thicker scars.
The researchers went on to make another critical finding. The mechanical stress of a fresh injury turns on a genetic program that leads to scar formation, and that program gets switched on through another protein called Yes-associated protein (YAP). When they blocked this protein with an existing eye drug called verteporfin, skin healed more slowly but without any hint of a scar.
It’s worth noting that scars aren’t just a cosmetic issue. Scars differ from unwounded skin in many ways. They lack hair follicles, glands that produce oil and sweat, and nerves for sensing pain or pressure. Because the fibers that make up scar tissue run parallel to each other instead of being more intricately interwoven, scars also lack the flexibility and strength of healthy skin.
These new findings therefore suggest it may one day be possible to allow wounds to heal without compromising the integrity of the skin. The findings also may have implications for many other medical afflictions that involve scarring, such as liver and lung fibrosis, burns, scleroderma, and scarring of heart tissue after a heart attack. That’s also quite a testament to sticking with a good postdoc project, wherever it may lead. One day, it may even improve public health!
References:
[1] Human skin wounds: A major and snowballing threat to public health and the economy. Sen CK, Gordillo GM, Roy S, Kirsner R, Lambert L, Hunt TK, Gottrup F, Gurtner GC, Longaker MT. Wound Repair Regen. 2009 Nov-Dec;17(6):763-771.
[2] Preventing Engrailed-1 activation in fibroblasts yields wound regeneration without scarring.
Mascharak S, desJardins-Park HE, Davitt MF, Griffin M, Borrelli MR, Moore AL, Chen K, Duoto B, Chinta M, Foster DS, Shen AH, Januszyk M, Kwon SH, Wernig G, Wan DC, Lorenz HP, Gurtner GC, Longaker MT. Science. 2021 Apr 23;372(6540):eaba2374.
[3] Skin fibrosis. Identification and isolation of a dermal lineage with intrinsic fibrogenic potential. Rinkevich Y, Walmsley GG, Hu MS, Maan ZN, Newman AM, Drukker M, Januszyk M, Krampitz GW, Gurtner GC, Lorenz HP, Weissman IL, Longaker MT. Science. 2015 Apr 17;348(6232):aaa2151.
Links:
Skin Health (National Institute of Arthritis and Musculoskeletal and Skin Diseases/NIH)
Scleroderma (NIAMS)
Michael Longaker (Stanford Medicine, Palo Alto, CA)
Geoffrey Gurtner (Stanford Medicine)
NIH Support: National Institute of General Medical Sciences; National Institute of Dental and Craniofacial Research
Study Finds Genetic Mutations in Healthy Human Tissues
Posted on by Dr. Francis Collins

The standard view of biology is that every normal cell copies its DNA instruction book with complete accuracy every time it divides. And thus, with a few exceptions like the immune system, cells in normal, healthy tissue continue to contain exactly the same genome sequence as was present in the initial single-cell embryo that gave rise to that individual. But new evidence suggests it may be time to revise that view.
By analyzing genetic information collected throughout the bodies of nearly 500 different individuals, researchers discovered that almost all had some seemingly healthy tissue that contained pockets of cells bearing particular genetic mutations. Some even harbored mutations in genes linked to cancer. The findings suggest that nearly all of us are walking around with genetic mutations within various parts of our bodies that, under certain circumstances, may have the potential to give rise to cancer or other health conditions.
Efforts such as NIH’s The Cancer Genome Atlas (TCGA) have extensively characterized the many molecular and genomic alterations underlying various types of cancer. But it has remained difficult to pinpoint the precise sequence of events that lead to cancer, and there are hints that so-called normal tissues, including blood and skin, might contain a surprising number of mutations —perhaps starting down a path that would eventually lead to trouble.
In the study published in Science, a team from the Broad Institute at MIT and Harvard, led by Gad Getz and postdoctoral fellow Keren Yizhak, along with colleagues from Massachusetts General Hospital, decided to take a closer look. They turned their attention to the NIH’s Genotype-Tissue Expression (GTEx) project.
The GTEx is a comprehensive public resource that shows how genes are expressed and controlled differently in various tissues throughout the body. To capture those important differences, GTEx researchers analyzed messenger RNA sequences within thousands of healthy tissue samples collected from people who died of causes other than cancer.
Getz, Yizhak, and colleagues wanted to use that extensive RNA data in another way: to detect mutations that had arisen in the DNA genomes of cells within those tissues. To do it, they devised a method for comparing those tissue-derived RNA samples to the matched normal DNA. They call the new method RNA-MuTect.
All told, the researchers analyzed RNA sequences from 29 tissues, including heart, stomach, pancreas, and fat, and matched DNA from 488 individuals in the GTEx database. Those analyses showed that the vast majority of people—a whopping 95 percent—had one or more tissues with pockets of cells carrying new genetic mutations.
While many of those genetic mutations are most likely harmless, some have known links to cancer. The data show that genetic mutations arise most often in the skin, esophagus, and lung tissues. This suggests that exposure to environmental elements—such as air pollution in the lung, carcinogenic dietary substances in the esophagus, or the ultraviolet radiation in sunlight that hits the skin—may play important roles in causing genetic mutations in different parts of the body.
The findings clearly show that, even within normal tissues, the DNA in the cells of our bodies isn’t perfectly identical. Rather, mutations constantly arise, and that makes our cells more of a mosaic of different mutational events. Sometimes those altered cells may have a subtle growth advantage, and thus continue dividing to form larger groups of cells with slightly changed genomic profiles. In other cases, those altered cells may remain in small numbers or perhaps even disappear.
It’s not yet clear to what extent such pockets of altered cells may put people at greater risk for developing cancer down the road. But the presence of these genetic mutations does have potentially important implications for early cancer detection. For instance, it may be difficult to distinguish mutations that are truly red flags for cancer from those that are harmless and part of a new idea of what’s “normal.”
To further explore such questions, it will be useful to study the evolution of normal mutations in healthy human tissues over time. It’s worth noting that so far, the researchers have only detected these mutations in large populations of cells. As the technology advances, it will be interesting to explore such questions at the higher resolution of single cells.
Getz’s team will continue to pursue such questions, in part via participation in the recently launched NIH Pre-Cancer Atlas. It is designed to explore and characterize pre-malignant human tumors comprehensively. While considerable progress has been made in studying cancer and other chronic diseases, it’s clear we still have much to learn about the origins and development of illness to build better tools for early detection and control.
Reference:
[1] RNA sequence analysis reveals macroscopic somatic clonal expansion across normal tissues. Yizhak K, Aguet F, Kim J, Hess JM, Kübler K, Grimsby J, Frazer R, Zhang H, Haradhvala NJ, Rosebrock D, Livitz D, Li X, Arich-Landkof E, Shoresh N, Stewart C, Segrè AV, Branton PA, Polak P, Ardlie KG, Getz G. Science. 2019 Jun 7;364(6444).
Links:
Genotype-Tissue Expression Program
The Cancer Genome Atlas (National Cancer Institute/NIH)
Pre-Cancer Atlas (National Cancer Institute/NIH)
Getz Lab (Broad Institute, Cambridge, MA)
NIH Support: Common Fund; National Heart, Lung, and Blood Institute; National Human Genome Research Institute; National Institute of Mental Health; National Cancer Institute; National Library of Medicine; National Institute on Drug Abuse; National Institute of Neurological Diseases and Stroke
Building a Smarter Bandage
Posted on by Dr. Francis Collins

Credit: Tufts University, Medford, MA
Smartphones, smartwatches, and smart electrocardiograms. How about a smart bandage?
This image features a prototype of a smart bandage equipped with temperature and pH sensors (lower right) printed directly onto the surface of a thin, flexible medical tape. You also see the “brains” of the operation: a microprocessor (upper left). When the sensors prompt the microprocessor, it heats up a hydrogel heating element in the bandage, releasing drugs and/or other healing substances on demand. It can also wirelessly transmit messages directly to a smartphone to keep patients and doctors updated.
While the smart bandage might help mend everyday cuts and scrapes, it was designed with the intent of helping people with hard-to-heal chronic wounds, such as leg and foot ulcers. Chronic wounds affect millions of Americans, including many seniors [1]. Such wounds are often treated at home and, if managed incorrectly, can lead to infections and potentially serious health problems.
Creative Minds: Taking Aim at Adverse Drug Reactions
Posted on by Dr. Francis Collins

Sherrie Divito
As a practicing dermatologist, Sherrie Divito sees lots of patients each week at Brigham and Women’s Hospital, Boston. She also sees lots of research opportunities. One that grabbed her attention is graft-versus-host disease (GvHD), which can arise after a bone-marrow transplant for leukemia, lymphoma, or various other diseases. What happens is immune cells in the donated marrow recognize a transplant patient’s body as “foreign” and launch an attack. Skin is often attacked first, producing a severe rash that is a harbinger of complications to come in other parts of the body.
But Divito saw something else: it’s virtually impossible to distinguish between an acute GvHD-caused rash and a severe skin reaction to drugs, from amoxicillin to carbamazepine. In her GvHD studies, Divito had been researching a recently identified class of immune cell called tissue-resident memory T (Trm) cells. They remain in skin rather than circulating in the bloodstream. The clinical similarities made Divito wonder whether Trm cells may also help to drive severe skin allergies to drugs.
Divito has received a 2016 NIH Director’s Early Independence Award to find out. If correct, Divito will help not only to improve the lives of thousands of people with GvHD, but potentially benefit the millions of other folks who experience adverse reactions to drug.
Next Page