technology
Can Bioprinted Skin Substitutes Replace Traditional Grafts for Treating Burn Injuries and Other Serious Skin Wounds?
Posted on by Lawrence Tabak, D.D.S., Ph.D.
Each year in the U.S., more than 500,000 people receive treatment for burn injuries and other serious skin wounds.1 To close the most severe wounds with less scarring, doctors often must surgically remove skin from one part of a person’s body and use it to patch the injured site. However, this is an intensive process, and some burn patients with extensive skin loss do not have sufficient skin available for grafting. Scientists have been exploring ways to repair these serious skin wounds without skin graft surgery.
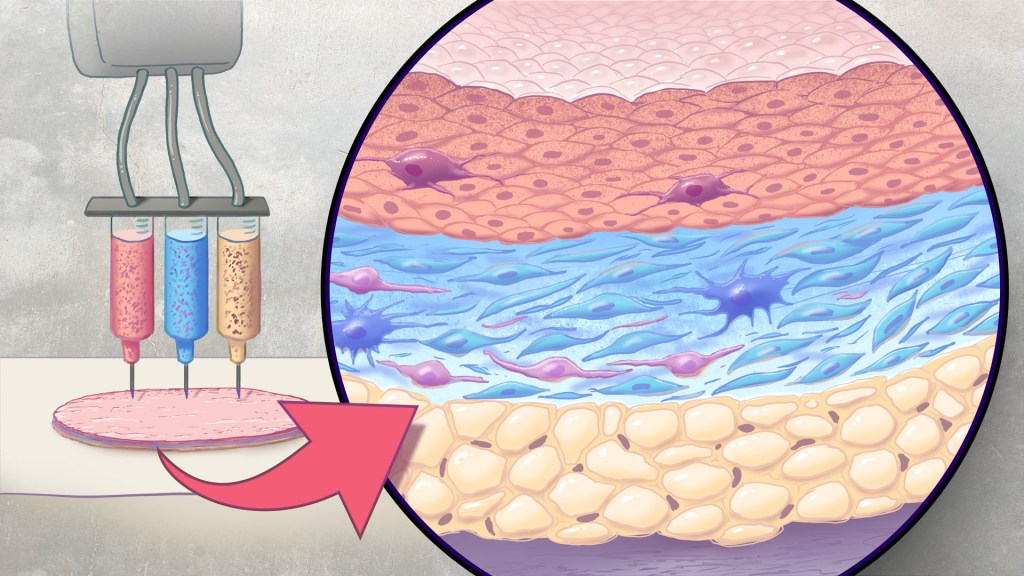
An NIH-funded team recently showed that bioprinted skin substitutes may serve as a promising alternative to traditional skin grafts in preclinical studies reported in Science Translational Medicine.2 The approach involves a portable skin bioprinter system that deposits multiple layers of skin directly into a wound. The recent findings add to evidence that bioprinting technology can successfully regenerate human-like skin to allow healing. While this approach has yet to be tested in people, it confirms that such technologies already can produce skin constructs with the complex structures and multiple cell types present in healthy human skin.
This latest work comes from a team led by Adam Jorgensen and Anthony Atala at Wake Forest School of Medicine’s Wake Forest Institute for Regenerative Medicine, Winston-Salem, NC. Members of the Atala lab and their colleagues had earlier shown it was possible to isolate two major skin cell types found in the skin’s outer (epidermis) and middle (dermis) layers from a small biopsy of healthy skin, expand the number of cells in the lab and then deliver the cells directly into an injury using a specially designed bioprinter.3 Using integrated imaging technology to scan a wound, computer software “prints” cells right into an injury, mimicking two of our skin’s three natural layers.
In the new study, Atala’s team has gone even further to construct skin substitutes that mimic the structure of human skin and that include six primary human skin cell types. They then used their bioprinter to produce skin constructs with all three layers found in healthy human skin: epidermis, dermis, and hypodermis.
To put their skin substitutes to the test, they first transplanted them into mice. Their studies showed that the bioprinted skin encouraged the rapid growth of new blood vessels and had other features of normal-looking, healthy skin. The researchers were able to confirm that their bioprinted skin implants successfully integrated into the animals’ regenerated skin to speed healing.
Studies in a pig model of wound healing added to evidence that such bioprinted implants can successfully repair full-thickness wounds, meaning those that extend through all three layers of skin. The bioprinted skin patches allowed for improved wound healing with less scarring. They also found that the bioprinted grafts encouraged activity in the skin from genes known to play important roles in wound healing.
It’s not yet clear if this approach will work as well in the clinic as it does in the lab. To make it feasible, the researchers note there’s a need for improved approaches to isolating and expanding the needed skin cell types. Nevertheless, these advances come as encouraging evidence that bioprinted skin substitutes could one day offer a promising alternative to traditional skin grafts with the capacity to help even more people with severe burns or other wounds.
References:
[1] Burn Incidence Fact Sheet. American Burn Association
[2] AM Jorgensen, et al. Multicellular bioprinted skin facilitates human-like skin architecture in vivo. Science Translational Medicine DOI: 10.1126/scitranslmed.adf7547 (2023).
[3] M Albanna, et al. In Situ Bioprinting of Autologous Skin Cells Accelerates Wound Healing of Extensive Excisional Full-Thickness Wounds. Scientific Reports DOI: 10.1038/s41598-018-38366-w (2019).
NIH Support: National Institute of Arthritis and Musculoskeletal and Skin Diseases
Rice-Sized Device Tests Brain Tumor’s Drug Responses During Surgery
Posted on by Lawrence Tabak, D.D.S., Ph.D.

Scientists have made remarkable progress in understanding the underlying changes that make cancer grow and have applied this knowledge to develop and guide targeted treatment approaches to vastly improve outcomes for people with many cancer types. And yet treatment progress for people with brain tumors known as gliomas—including the most aggressive glioblastomas—has remained slow. One reason is that doctors lack tests that reliably predict which among many therapeutic options will work best for a given tumor.
Now an NIH-funded team has developed a miniature device with the potential to change this for the approximately 25,000 people diagnosed with brain cancers in the U.S. each year [1]. When implanted into cancerous brain tissue during surgery, the rice-sized drug-releasing device can simultaneously conduct experiments to measure a tumor’s response to more than a dozen drugs or drug combinations. What’s more, a small clinical trial reported in Science Translational Medicine offers the first evidence in people with gliomas that these devices can safely offer unprecedented insight into tumor-specific drug responses [2].
These latest findings come from a Brigham and Women’s Hospital, Boston, team led by Pierpaolo Peruzzi and Oliver Jonas. They recognized that drug-screening studies conducted in cells or tissue samples in the lab too often failed to match what happens in people with gliomas undergoing cancer treatment. Wide variation within individual brain tumors also makes it hard to predict a tumor’s likely response to various treatment options.
It led them to an intriguing idea: Why not test various therapeutic options in each patient’s tumor? To do it, they developed a device, about six millimeters long, that can be inserted into a brain tumor during surgery to deliver tiny doses of up to 20 drugs. Doctors can then remove and examine the drug-exposed cancerous tissue in the laboratory to determine each treatment’s effects. The data can then be used to guide subsequent treatment decisions, according to the researchers.
In the current study, the researchers tested their device on six study volunteers undergoing brain surgery to remove a glioma tumor. For each volunteer, the device was implanted into the tumor and remained in place for about two to three hours while surgeons worked to remove most of the tumor. Next, the device was taken out along with the last piece of a tumor at the end of the surgery for further study of drug responses.
Importantly, none of the study participants experienced any adverse effects from the device. Using the devices, the researchers collected valuable data, including how a tumor’s response changed with varying drug concentrations or how each treatment led to molecular changes in the cancerous cells.
More research is needed to better understand how use of such a device might change treatment and patient outcomes in the longer term. The researchers note that it would take more than a couple of hours to determine how treatments produce less immediate changes, such as immune responses. As such, they’re now conducting a follow-up trial to test a possible two-stage procedure, in which their device is inserted first using minimally invasive surgery 72 hours prior to a planned surgery, allowing longer exposure of tumor tissue to drugs prior to a tumor’s surgical removal.
Many questions remain as they continue to optimize this approach. However, it’s clear that such a device gives new meaning to personalized cancer treatment, with great potential to improve outcomes for people living with hard-to-treat gliomas.
References:
[1] National Cancer Institute Surveillance, Epidemiology, and End Results Program. Cancer Stat Facts: Brain and Other Nervous System Cancer.
[2] Peruzzi P et al. Intratumoral drug-releasing microdevices allow in situ high-throughput pharmaco phenotyping in patients with gliomas. Science Translational Medicine DOI: 10.1126/scitranslmed.adi0069 (2023).
Links:
Brain Tumors – Patient Version (National Cancer Institute/NIH)
Pierpaolo Peruzzi (Brigham and Women’s Hospital, Boston, MA)
Jonas Lab (Brigham and Women’s Hospital, Boston, MA)
NIH Support: National Cancer Institute, National Institute of Biomedical Imaging and Bioengineering, National Institute of Neurological Disorders and Stroke
New Approach to ‘Liquid Biopsy’ Relies on Repetitive RNA in the Bloodstream
Posted on by Lawrence Tabak, D.D.S., Ph.D.

It’s always best to diagnose cancer at an early stage when treatment is most likely to succeed. Unfortunately, far too many cancers are still detected only after cancer cells have escaped from a primary tumor and spread to distant parts of the body. This explains why there’s been so much effort in recent years to develop liquid biopsies, which are tests that can pick up on circulating cancer cells or molecular signs of cancer in blood or other bodily fluids and reliably trace them back to the organ in which a potentially life-threatening tumor is growing.
Earlier methods to develop liquid biopsies for detecting cancers often have relied on the presence of cancer-related proteins and/or DNA in the bloodstream. Now, an NIH-supported research team has encouraging evidence to suggest that this general approach to detecting cancers—including aggressive pancreatic cancers—may work even better by taking advantage of signals from a lesser-known form of genetic material called noncoding RNA.
The findings reported in Nature Biomedical Engineering suggest that the new liquid biopsy approach may aid in the diagnosis of many forms of cancer [1]. The studies show that the sensitivity of the tests varies—a highly sensitive test is one that rarely misses cases of disease. However, they already have evidence that millions of circulating RNA molecules may hold promise for detecting cancers of the liver, esophagus, colon, stomach, and lung.
How does it work? The human genome contains about 3 billion paired DNA letters. Most of those letters are transcribed, or copied, into single-stranded RNA molecules. While RNA is best known for encoding proteins that do the work of the cell, most RNA never gets translated into proteins at all. This noncoding RNA includes repetitive RNA that can be transcribed from millions of repeat elements—patterns of the same few DNA letters occurring multiple times in the genome.
Common approaches to studying RNA don’t analyze repetitive RNA, so its usefulness as a diagnostic tool has been unclear—until recently. Last year, the lab of Daniel Kim at the University of California, Santa Cruz reported [2] that a key genetic mutation that occurs early on in some cancers causes repetitive RNA molecules to be secreted in large quantities from cancer cells, even at the earliest stages of cancer. Non-cancerous cells, by comparison, release much less repetitive RNA.
The findings suggested that liquid biopsy tests that look for this repetitive, noncoding RNA might offer a powerful new way to detect cancers sooner, according to the authors. But first they needed a method capable of measuring it. Due to its oftentimes uncertain functions, the researchers have referred to repetitive, noncoding RNA as “dark matter.”
Using a liquid biopsy platform they developed called COMPLETE-seq, Kim’s team trained computers to detect cancers by looking for patterns in RNA data. The platform enables sequencing and analysis of all protein coding and noncoding RNAs—including any RNA from more than 5 million repeat elements—present in a blood sample. They found that their classifiers worked better when repetitive RNAs were included. The findings lend support to the idea that repetitive, noncoding RNA in the bloodstream is a rich source of information for detecting cancers, which has previously been overlooked.
In a study comparing blood samples from healthy people to those with pancreatic cancer, the COMPLETE-seq technology showed that nearly all people in the study with pancreatic cancer had more repetitive, noncoding RNA in their blood samples compared to healthy people, according to the researchers. They used the COMPLETE-seq test on blood samples from people with other types of cancer as well. For example, their test accurately detected 91% of colorectal cancer samples and 93% of lung cancer samples.
They now plan to look at many more cancer types with samples from additional patients representing a broad range of cancer stages. The goal is to develop a single RNA liquid biopsy test that could detect multiple forms of cancer with a high degree of accuracy and specificity. They note that such a test might also be used to guide treatment decisions and more readily detect a cancer’s recurrence. The hope is that one day a comprehensive liquid biopsy test including coding and noncoding RNA will catch many more cancers sooner, when treatment can be most successful.
References:
[1] RE Reggiardo et al. Profiling of repetitive RNA sequences in the blood plasma of patients with cancer. Nature Biomedical Engineering DOI: 10.1038/s41551-023-01081-7 (2023).
[2] RE Reggiardo et al. Mutant KRAS regulates transposable element RNA and innate immunity via KRAB zinc-finger genes. Cell Reports DOI: 10.1016/j.celrep.2022.111104 (2022).
Links:
Daniel Kim Lab (UC Santa Cruz)
Cancer Screening Overview (National Cancer Institute/NIH)
Early Detection (National Cancer Institute/NIH)
NIH Support: National Cancer Institute, National Heart, Lung, and Blood Institute, National Institute of Diabetes and Digestive and Kidney Diseases
Mapping Immune Cell “Neighborhoods” in Psoriasis to Understand its Course
Posted on by Lawrence Tabak, D.D.S., Ph.D.

“Location, location, location.” While most of us know this phrase as a real estate adage, location—specifically that of various cell types—is becoming a key area of investigation in studying human disease. New techniques are enabling scientists to understand where certain cells are with respect to one another and how changes in their activity may affect your overall health.
In one recent example of the power of this approach, NIH-funded researchers [1] used a sophisticated method to map immune cells within human skin to get a more detailed picture of psoriasis, a common, chronic disease in which the immune system becomes overactive leading to skin inflammation. People with psoriasis develop patches of itchy, red, and flaky lesions on their skin, which can be mild to severe. For reasons that aren’t entirely clear, they’re also at higher risk for developing a wide range of other health conditions, including a unique form of arthritis known as psoriatic arthritis, diabetes, mental health issues, heart problems, and more.
The hope is that these newly drawn, precise maps of cellular “neighborhoods” in human skin will help chart the precise course of this disease to understand better the differences between mild and more severe forms. They may also yield important clues as to why people with psoriasis develop other health problems more often than people without psoriasis.
In the new study, a team including Jose Scher and Shruti Naik, NYU Langone, New York, analyzed immune cells within 25 skin samples from 14 volunteers, including those with active psoriasis, those with psoriasis but no active lesions, and people with healthy skin who do not have psoriasis. The researchers relied on a sophisticated approach called spatial transcriptomics [2] to map out what happens at the single-cell level within the samples.
In earlier approaches to single-cell analysis, researchers first would separate cells from the tissue they came from. While they could measure gene activity within those cells at the individual level, they couldn’t put things back together to see how they all fit. With spatial transcriptomics, it’s now possible to molecularly profile single cells to measure their activity in a tissue sample while also mapping their locations with respect to other cells.
The new study led to some intriguing findings. For instance, certain immune cells, specifically B cells, moved to the upper layers of the skin during active disease. That’s notable because prior studies had been unable to capture B cells in the skin adequately, and these cells are thought to play an important role in the disease.
Interestingly, the spatial cellular maps revealed inflammatory regions in both actively inflamed skin and in skin that appeared healthy. This finding highlights the fact that the inflammation that goes with psoriasis can affect the skin, and likely other parts of the body, in ways that aren’t easily observed. In future studies, the researchers want to explore how the presence of psoriasis and its underlying changes in immune cell activity may influence other organs and tissues beneath the skin.
Their fine-scale maps also showed increased gene activity in dozens of molecular pathways that are tied to metabolism and the control of lipid levels. That’s especially interesting because these factors are known to go awry in diabetes and heart conditions, which happen more often in people with psoriasis compared to those without. They also could see in their maps that this altered activity sometimes occurred in clear skin distant from any apparent lesions.
Having discovered such signals with potential consequences for other parts of the body, the researchers report that they’re working to understand how inflammatory immune cells and processes in the skin may lead to more widespread disease processes that affect other parts of the body. They plan to conduct similar studies in larger groups of people with and without active psoriasis lesions and studies following individuals with psoriasis over time. They’ll also explore questions about why people respond differently to the same anti-inflammatory treatment regimens.
To speed the process of discovery, they’ve made their maps and associated data freely available as a resource for the scientific community. About 7.5 million adults in the U.S. and millions more worldwide have psoriasis and associated psoriatic conditions [3]. The hope is that these maps will one day help to steer them toward a healthier future.
References:
[1] Spatial transcriptomics stratifies psoriatic disease severity by emergent cellular ecosystems. Castillo RL, Sidhu I, Dolgalev I, Chu T, Prystupa A, Subudhi I, Yan D, Konieczny P, Hsieh B, Haberman RH, Selvaraj S, Shiomi T, Medina R, Girija PV, Heguy A, Loomis CA, Chiriboga L, Ritchlin C, Garcia-Hernandez ML, Carucci J, Meehan SA, Neimann AL, Gudjonsson JE, Scher JU, Naik S. Sci Immunol. 2023 Jun 8;8(84):eabq7991. doi: 10.1126/sciimmunol.abq7991.
[2] Method of the Year: spatially resolved transcriptomics. Marx V. Nat Methods. 2021 Jan;18(1):9-14. doi: 10.1038/s41592-020-01033-y.
[3] Psoriasis Prevalence in Adults in the United States. Armstrong AW, Mehta MD, Schupp CW, Gondo GC, Bell SJ, Griffiths CEM. JAMA Dermatol. 2021 Aug 1;157(8):940-946. doi: 10.1001/jamadermatol.2021.2007.
Links:
Psoriasis (National Institute of Arthritis and Musculoskeletal and Skin Diseases/NIH)
Jose Scher (NYU Langone Health, New York, NY)
Shruti Naik (NYU Langone Health, New York, NY)
NIH Support: National Cancer Institute, National Institute of Arthritis and Musculoskeletal and Skin Diseases, National Center for Advancing Translational Sciences, National Institute of Allergy and Infectious Diseases
Wearable Scanner Tracks Brain Activity While Body Moves
Posted on by Dr. Francis Collins
Credit: Wellcome Centre for Human Neuroimaging, University College London.
In recent years, researchers fueled by the BRAIN Initiative and many other NIH-supported efforts have made remarkable progress in mapping the human brain in all its amazing complexity. Now, a powerful new imaging technology promises to further transform our understanding [1]. This wearable scanner, for the first time, enables researchers to track neural activity in people in real-time as they do ordinary things—be it drinking tea, typing on a keyboard, talking to a friend, or even playing paddle ball.
This new so-called magnetoencephalography (MEG) brain scanner, which looks like a futuristic cross between a helmet and a hockey mask, is equipped with specialized “quantum” sensors. When placed directly on the scalp surface, these new MEG scanners can detect weak magnetic fields generated by electrical activity in the brain. While current brain scanners weigh in at nearly 1,000 pounds and require people to come to a special facility and remain absolutely still, the new system weighs less than 2 pounds and is capable of generating 3D images even when a person is making motions.
Basic Research: Building a Firm Foundation for Biomedicine
Posted on by Dr. Francis Collins

Credit: National Institute of Allergy and Infectious Diseases, NIH
A major part of NIH’s mission is to support basic research that generates fundamental knowledge about the nature and behavior of living systems. Such knowledge serves as the foundation for the biomedical advances needed to protect and improve our health—and the health of generations to come.
Of course, it’s often hard to predict how this kind of basic research might benefit human populations, and the lag time between discovery and medical application (if that happens at all) can be quite long. Some might argue, therefore, that basic research is not a good use of funds, and all of NIH’s support should go to specific disease targets.
To counter that perception, I’m pleased to share some new findings that underscore the importance of publicly supported basic research. In an analysis of more than 28 million papers in the PubMed.gov database, researchers found NIH contributed to published research that was associated with every single one of the 210 new drugs approved by the Food and Drug Administration from 2010 through 2016 [1]. More than 90 percent of that contributory research was basic—that is, related to the discovery of fundamental biological mechanisms, rather than actual development of the drugs themselves.
Powerful Antibiotics Found in Dirt
Posted on by Dr. Francis Collins

Caption: Researchers found a new class of antibiotics in a collection of about 2,000 soil samples.
Credit: Sean Brady, The Rockefeller University, New York City
Many of us think of soil as lifeless dirt. But, in fact, soil is teeming with a rich array of life: microbial life. And some of those tiny, dirt-dwelling microorganisms—bacteria that produce antibiotic compounds that are highly toxic to other bacteria—may provide us with valuable leads for developing the new drugs we so urgently need to fight antibiotic-resistant infections.
Recently, NIH-funded researchers discovered a new class of antibiotics, called malacidins, by analyzing the DNA of the bacteria living in more than 2,000 soil samples, including many sent by citizen scientists living all across the United States [1]. While more work is needed before malacidins can be tried in humans, the compounds successfully killed several types of multidrug-resistant bacteria in laboratory tests. Most impressive was the ability of malacadins to wipe out methicillin-resistant Staphylococcus aureus (MRSA) skin infections in rats. Often referred to as a “super bug,” MRSA threatens the lives of tens of thousands of Americans each year [2].
Sequencing Human Genome with Pocket-Sized “Nanopore” Device
Posted on by Dr. Francis Collins

Caption: MinION sequencing device plugged into a laptop/Oxford Nanopore Technologies
It’s hard to believe, but it’s been almost 15 years since we successfully completed the Human Genome Project, ahead of schedule and under budget. I was proud to stand with my international colleagues in a celebration at the Library of Congress on April 14, 2003 (which happens to be my birthday), to announce that we had stitched together the very first reference sequence of the human genome at a total cost of about $400 million. As remarkable as that achievement was, it was just the beginning of our ongoing effort to understand the human genome, and to use that understanding to improve human health.
That first reference human genome was sequenced using automated machines that were the size of small phone booths. Since then, breathtaking progress has been made in developing innovative technologies that have made DNA sequencing far easier, faster, and more affordable. Now, a report in Nature Biotechnology highlights the latest advance: the sequencing and assembly of a human genome using a pocket-sized device [1]. It was generated using several “nanopore” devices that can be purchased online with a “starter kit” for just $1,000. In fact, this new genome sequence—completed in a matter of weeks—includes some notoriously hard-to-sequence stretches of DNA, filling several key gaps in our original reference genome.
Next Page