DNA sequencing
All of Us: Release of Nearly 100,000 Whole Genome Sequences Sets Stage for New Discoveries
Posted on by Joshua Denny, M.D., M.S., and Lawrence Tabak, D.D.S., Ph.D.

Nearly four years ago, NIH opened national enrollment for the All of Us Research Program. This historic program is building a vital research community within the United States of at least 1 million participant partners from all backgrounds. Its unifying goal is to advance precision medicine, an emerging form of health care tailored specifically to the individual, not the average patient as is now often the case. As part of this historic effort, many participants have offered DNA samples for whole genome sequencing, which provides information about almost all of an individual’s genetic makeup.
Earlier this month, the All of Us Research Program hit an important milestone. We released the first set of nearly 100,000 whole genome sequences from our participant partners. The sequences are stored in the All of Us Researcher Workbench, a powerful, cloud-based analytics platform that makes these data broadly accessible to registered researchers.
The All of Us Research Program and its many participant partners are leading the way toward more equitable representation in medical research. About half of this new genomic information comes from people who self-identify with a racial or ethnic minority group. That’s extremely important because, until now, over 90 percent of participants in large genomic studies were of European descent. This lack of diversity has had huge impacts—deepening health disparities and hindering scientific discovery from fully benefiting everyone.
The Researcher Workbench also contains information from many of the participants’ electronic health records, Fitbit devices, and survey responses. Another neat feature is that the platform links to data from the U.S. Census Bureau’s American Community Survey to provide more details about the communities where participants live.
This unique and comprehensive combination of data will be key in transforming our understanding of health and disease. For example, given the vast amount of data and diversity in the Researcher Workbench, new diseases are undoubtedly waiting to be uncovered and defined. Many new genetic variants are also waiting to be identified that may better predict disease risk and response to treatment.
To speed up the discovery process, these data are being made available, both widely and wisely. To protect participants’ privacy, the program has removed all direct identifiers from the data and upholds strict requirements for researchers seeking access. Already, more than 1,500 scientists across the United States have gained access to the Researcher Workbench through their institutions after completing training and agreeing to the program’s strict rules for responsible use. Some of these researchers are already making discoveries that promote precision medicine, such as finding ways to predict how to best to prevent vision loss in patients with glaucoma.
Beyond making genomic data available for research, All of Us participants have the opportunity to receive their personal DNA results, at no cost to them. So far, the program has offered genetic ancestry and trait results to more than 100,000 participants. Plans are underway to begin sharing health-related DNA results on hereditary disease risk and medication-gene interactions later this year.
This first release of genomic data is a huge milestone for the program and for health research more broadly, but it’s also just the start. The program’s genome centers continue to generate the genomic data and process about 5,000 additional participant DNA samples every week.
The ultimate goal is to gather health data from at least 1 million or more people living in the United States, and there’s plenty of time to join the effort. Whether you would like to contribute your own DNA and health information, engage in research, or support the All of Us Research Program as a partner, it’s easy to get involved. By taking part in this historic program, you can help to build a better and more equitable future for health research and precision medicine.
Note: Joshua Denny, M.D., M.S., is the Chief Executive Officer of NIH’s All of Us Research Program.
Links:
All of Us Research Program (NIH)
Join All of Us (NIH)
All of Us: Partnering Together for the Future of Precision Medicine
Posted on by Dr. Francis Collins

Over the past year, it’s been so inspiring to watch tens of thousands of people across the country selflessly step forward for vaccine trials and other research studies to combat COVID-19. And they are not alone. Many generous folks are volunteering to take part in other types of NIH-funded research that will improve health all across the spectrum, including the more than 360,000 who’ve already enrolled in the pioneering All of Us Research Program.
Now in its second year, All of Us is building a research community of 1 million participant partners to help us learn more about how genetics, environment, and lifestyle interact to influence disease and affect health. So far, more than 80 percent of participants who have completed all the initial enrollment steps are Black, Latino, rural, or from other communities historically underrepresented in biomedical research.
This community will build a diverse foundation for precision medicine, in which care is tailored to the individual, not the average patient as is now often the case. What’s also paradigm shifting about All of Us is its core value of sharing information back with participants about themselves. It is all done responsibly through each participant’s personal All of Us online account and with an emphasis on protecting privacy.
All of Us participants share their health information in many ways, such as taking part in surveys, offering access to their electronic health records, and providing biosamples (blood, urine, and/or saliva). In fact, researchers recently began genotyping and sequencing the DNA in some of those biosamples, and then returning results from analyses to participants who’ve indicated they’d like to receive such information. This first phase of genotyping DNA analysis will provide insights into their genetic ancestry and four traits, including bitter taste perception and tolerance for lactose.
Results of a second sequencing phase of DNA analysis will likely be ready in the coming year. These personalized reports will give interested participants information about how their bodies are likely to react to certain medications and about whether they face an increased risk of developing certain health conditions, such as some types of cancer or heart disease. To help participants better understand the results, they can make a phone appointment with a genetic counselor who is affiliated with the program.
This week, I had the pleasure of delivering the keynote address at the All of Us Virtual Face-to-Face. This lively meeting was attended by a consortium of more than 2,000 All of Us senior staff, program leads with participating healthcare provider organizations and federally qualified health centers, All of Us-supported researchers, community partners, and the all-important participant ambassadors.
If you are interested in becoming part of the All of Us community, I welcome you—there’s plenty of time to get involved! To learn more, just go to Join All of Us.
Links:
All of Us Research Program (NIH)
Join All of Us (NIH)
Whole-Genome Sequencing Plus AI Yields Same-Day Genetic Diagnoses
Posted on by Dr. Francis Collins

Back in April 2003, when the international Human Genome Project successfully completed the first reference sequence of the human DNA blueprint, we were thrilled to have achieved that feat in just 13 years. Sure, the U.S. contribution to that first human reference sequence cost an estimated $400 million, but we knew (or at least we hoped) that the costs would come down quickly, and the speed would accelerate. How far we’ve come since then! A new study shows that whole genome sequencing—combined with artificial intelligence (AI)—can now be used to diagnose genetic diseases in seriously ill babies in less than 24 hours.
Take a moment to absorb this. I would submit that there is no other technology in the history of planet Earth that has experienced this degree of progress in speed and affordability. And, at the same time, DNA sequence technology has achieved spectacularly high levels of accuracy. The time-honored adage that you can only get two out of three for “faster, better, and cheaper” has been broken—all three have been dramatically enhanced by the advances of the last 16 years.
Rapid diagnosis is critical for infants born with mysterious conditions because it enables them to receive potentially life-saving interventions as soon as possible after birth. In a study in Science Translational Medicine, NIH-funded researchers describe development of a highly automated, genome-sequencing pipeline that’s capable of routinely delivering a diagnosis to anxious parents and health-care professionals dramatically earlier than typically has been possible [1].
While the cost of rapid DNA sequencing continues to fall, challenges remain in utilizing this valuable tool to make quick diagnostic decisions. In most clinical settings, the wait for whole-genome sequencing results still runs more than two weeks. Attempts to obtain faster results also have been labor intensive, requiring dedicated teams of experts to sift through the data, one sample at a time.
In the new study, a research team led by Stephen Kingsmore, Rady Children’s Institute for Genomic Medicine, San Diego, CA, describes a streamlined approach that accelerates every step in the process, making it possible to obtain whole-genome test results in a median time of about 20 hours and with much less manual labor. They propose that the system could deliver answers for 30 patients per week using a single genome sequencing instrument.
Here’s how it works: Instead of manually preparing blood samples, his team used special microbeads to isolate DNA much more rapidly with very little labor. The approach reduced the time for sample preparation from 10 hours to less than three. Then, using a state-of-the-art DNA sequencer, they sequence those samples to obtain good quality whole genome data in just 15.5 hours.
The next potentially time-consuming challenge is making sense of all that data. To speed up the analysis, Kingsmore’s team took advantage of a machine-learning system called MOON. The automated platform sifts through all the data using artificial intelligence to search for potentially disease-causing variants.
The researchers paired MOON with a clinical language processing system, which allowed them to extract relevant information from the child’s electronic health records within seconds. Teaming that patient-specific information with data on more than 13,000 known genetic diseases in the scientific literature, the machine-learning system could pick out a likely disease-causing mutation out of 4.5 million potential variants in an impressive 5 minutes or less!
To put the system to the test, the researchers first evaluated its ability to reach a correct diagnosis in a sample of 101 children with 105 previously diagnosed genetic diseases. In nearly every case, the automated diagnosis matched the opinions reached previously via the more lengthy and laborious manual interpretation of experts.
Next, the researchers tested the automated system in assisting diagnosis of seven seriously ill infants in the intensive care unit, and three previously diagnosed infants. They showed that their automated system could reach a diagnosis in less than 20 hours. That’s compared to the fastest manual approach, which typically took about 48 hours. The automated system also required about 90 percent less manpower.
The system nailed a rapid diagnosis for 3 of 7 infants without returning any false-positive results. Those diagnoses were made with an average time savings of more than 22 hours. In each case, the early diagnosis immediately influenced the treatment those children received. That’s key given that, for young children suffering from serious and unexplained symptoms such as seizures, metabolic abnormalities, or immunodeficiencies, time is of the essence.
Of course, artificial intelligence may never replace doctors and other healthcare providers. Kingsmore notes that 106 years after the invention of the autopilot, two pilots are still required to fly a commercial aircraft. Likewise, health care decisions based on genome interpretation also will continue to require the expertise of skilled physicians.
Still, such a rapid automated system will prove incredibly useful. For instance, this system can provide immediate provisional diagnosis, allowing the experts to focus their attention on more difficult unsolved cases or other needs. It may also prove useful in re-evaluating the evidence in the many cases in which manual interpretation by experts fails to provide an answer.
The automated system may also be useful for periodically reanalyzing data in the many cases that remain unsolved. Keeping up with such reanalysis is a particular challenge considering that researchers continue to discover hundreds of disease-associated genes and thousands of variants each and every year. The hope is that in the years ahead, the combination of whole genome sequencing, artificial intelligence, and expert care will make all the difference in the lives of many more seriously ill babies and their families.
Reference:
[1] Diagnosis of genetic diseases in seriously ill children by rapid whole-genome sequencing and automated phenotyping and interpretation. Clark MM, Hildreth A, Batalov S, Ding Y, Chowdhury S, Watkins K, Ellsworth K, Camp B, Kint CI, Yacoubian C, Farnaes L, Bainbridge MN, Beebe C, Braun JJA, Bray M, Carroll J, Cakici JA, Caylor SA, Clarke C, Creed MP, Friedman J, Frith A, Gain R, Gaughran M, George S, Gilmer S, Gleeson J, Gore J, Grunenwald H, Hovey RL, Janes ML, Lin K, McDonagh PD, McBride K, Mulrooney P, Nahas S, Oh D, Oriol A, Puckett L, Rady Z, Reese MG, Ryu J, Salz L, Sanford E, Stewart L, Sweeney N, Tokita M, Van Der Kraan L, White S, Wigby K, Williams B, Wong T, Wright MS, Yamada C, Schols P, Reynders J, Hall K, Dimmock D, Veeraraghavan N, Defay T, Kingsmore SF. Sci Transl Med. 2019 Apr 24;11(489).
Links:
DNA Sequencing Fact Sheet (National Human Genome Research Institute/NIH)
Genomics and Medicine (NHGRI/NIH)
Genetic and Rare Disease Information Center (National Center for Advancing Translational Sciences/NIH)
Stephen Kingsmore (Rady Children’s Institute for Genomic Medicine, San Diego, CA)
NIH Support: National Institute of Child Health and Human Development; National Human Genome Research Institute; National Center for Advancing Translational Sciences
Study Shows Genes Unique to Humans Tied to Bigger Brains
Posted on by Dr. Francis Collins
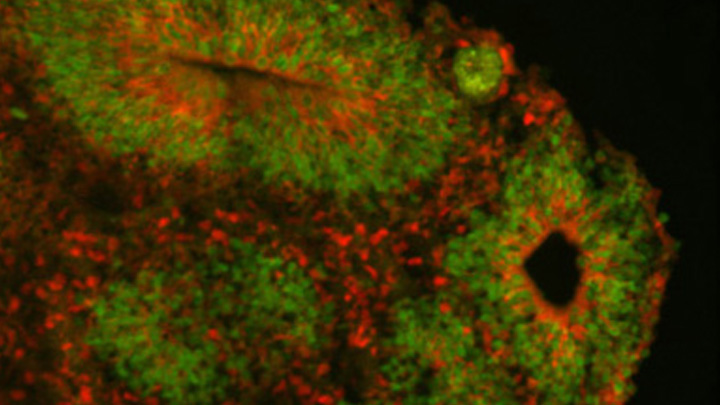
Caption: Cortical organoid, showing radial glial stem cells (green) and cortical neurons (red).
Credit: Sofie Salama, University of California, Santa Cruz
In seeking the biological answer to the question of what it means to be human, the brain’s cerebral cortex is a good place to start. This densely folded, outer layer of grey matter, which is vastly larger in Homo sapiens than in other primates, plays an essential role in human consciousness, language, and reasoning.
Now, an NIH-funded team has pinpointed a key set of genes—found only in humans—that may help explain why our species possesses such a large cerebral cortex. Experimental evidence shows these genes prolong the development of stem cells that generate neurons in the cerebral cortex, which in turn enables the human brain to produce more mature cortical neurons and, thus, build a bigger cerebral cortex than our fellow primates.
That sounds like a great advantage for humans! But there’s a downside. Researchers found the same genomic changes that facilitated the expansion of the human cortex may also render our species more susceptible to certain rare neurodevelopmental disorders.
Sequencing Human Genome with Pocket-Sized “Nanopore” Device
Posted on by Dr. Francis Collins

Caption: MinION sequencing device plugged into a laptop/Oxford Nanopore Technologies
It’s hard to believe, but it’s been almost 15 years since we successfully completed the Human Genome Project, ahead of schedule and under budget. I was proud to stand with my international colleagues in a celebration at the Library of Congress on April 14, 2003 (which happens to be my birthday), to announce that we had stitched together the very first reference sequence of the human genome at a total cost of about $400 million. As remarkable as that achievement was, it was just the beginning of our ongoing effort to understand the human genome, and to use that understanding to improve human health.
That first reference human genome was sequenced using automated machines that were the size of small phone booths. Since then, breathtaking progress has been made in developing innovative technologies that have made DNA sequencing far easier, faster, and more affordable. Now, a report in Nature Biotechnology highlights the latest advance: the sequencing and assembly of a human genome using a pocket-sized device [1]. It was generated using several “nanopore” devices that can be purchased online with a “starter kit” for just $1,000. In fact, this new genome sequence—completed in a matter of weeks—includes some notoriously hard-to-sequence stretches of DNA, filling several key gaps in our original reference genome.
Studies of Dogs, Mice, and People Provide Clues to OCD
Posted on by Dr. Francis Collins

Thinkstock/wildpixel
Chances are you know someone with obsessive-compulsive disorder (OCD). It’s estimated that more than 2 million Americans struggle with this mental health condition, characterized by unwanted recurring thoughts and/or repetitive behaviors, such as excessive hand washing or constant counting of objects. While we know that OCD tends to run in families, it’s been frustratingly difficult to identify specific genes that influence OCD risk.
Now, an international research team, partly funded by NIH, has made progress thanks to an innovative genomic approach involving dogs, mice, and people. The strategy allowed them to uncover four genes involved in OCD that turn out to play a role in synapses, where nerve impulses are transmitted between neurons in the brain. While more research is needed to confirm the findings and better understand the molecular mechanisms of OCD, these findings offer important new leads that could point the way to more effective treatments.
Happy New Year: Looking Back at 2016 Research Highlights
Posted on by Dr. Francis Collins
 Happy New Year! While everyone was busy getting ready for the holidays, the journal Science announced its annual compendium of scientific Breakthroughs of the Year. If you missed it, the winner for 2016 was the detection of gravitational waves—tiny ripples in the fabric of spacetime created by the collision of two black holes 1.3 billion years ago! It’s an incredible discovery, and one that Albert Einstein predicted a century ago.
Happy New Year! While everyone was busy getting ready for the holidays, the journal Science announced its annual compendium of scientific Breakthroughs of the Year. If you missed it, the winner for 2016 was the detection of gravitational waves—tiny ripples in the fabric of spacetime created by the collision of two black holes 1.3 billion years ago! It’s an incredible discovery, and one that Albert Einstein predicted a century ago.
Among the nine other advances that made the first cut for Breakthrough of the Year, several involved the biomedical sciences. As I’ve done in previous years (here and here), I’ll kick off this New Year by taking a quick look of some of the breakthroughs that directly involved NIH support:
Genome Sequencing: Exploring the Diagnostic Promise
Posted on by Dr. Francis Collins
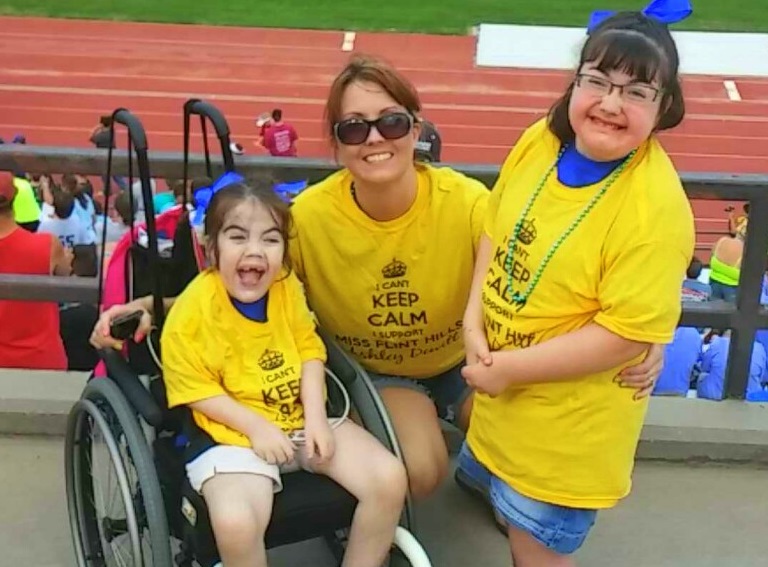

Caption: Whole genome sequencing revealed that sisters Addison and Trinity Hanners, ages 7 and 10, shown here with their mother Hanna, have a rare syndrome caused by a mutation in the MAGEL2 gene.
Credit: Courtesy of the Hanners family
At the time that we completed a draft of the 3 billion letters of the human genome about a decade ago, it would have cost about $100 million to sequence a second human genome. Today, thanks to advances in DNA sequencing technology, it will soon be possible to sequence your genome or mine for $1,000 or less. All of this progress has made genome sequencing a far more realistic clinical option to consider for people, especially children, who suffer from baffling disorders that can’t be precisely diagnosed by other medical tests.
While researchers are still in the process of evaluating genome sequencing for routine clinical use, and data analysis continues to be a major challenge, one area of considerable promise centers on neurodevelopmental disorders. Such disorders—which affect about 3 percent of children—range from relatively common conditions like autism spectrum disorder to very rare conditions that impair the development of the brain or central nervous system. In the latest study, an NIH-funded research team reports that sequencing either a patient’s whole genome or whole exome (the 1.5 percent of the genome that encodes proteins) appears to be an effective—as well as a cost-effective—strategy for diagnosing neurodevelopmental disorders that have eluded diagnosis through standard means.
Next Page
 We humans are wired to search for a causative agent when something bad happens. When someone develops cancer, we seek a reason. Maybe cancer runs in the family. Or perhaps the person smoked, never wore sunscreen, or drank too much alcohol. At some level, those are reasonable assumptions, as genes, lifestyle, and environment do play important roles in cancer. But a new study claims that the reason why many people get cancer is simply just bad luck.
We humans are wired to search for a causative agent when something bad happens. When someone develops cancer, we seek a reason. Maybe cancer runs in the family. Or perhaps the person smoked, never wore sunscreen, or drank too much alcohol. At some level, those are reasonable assumptions, as genes, lifestyle, and environment do play important roles in cancer. But a new study claims that the reason why many people get cancer is simply just bad luck.