blood lipids
Protein Links Gut Microbes, Biological Clocks, and Weight Gain
Posted on by Dr. Francis Collins

Caption: Lipids (red) inside mouse intestinal cells with and without NFIL3.
Credit: Lora V. Hooper, University of Texas Southwestern Medical Center, Dallas
The American epidemic of obesity is a major public health concern, and keeping off the extra pounds is a concern for many of us. Yet it can also be a real challenge for people who may eat normally but get their days and nights mixed up, including night-shift workers and those who regularly travel overseas. Why is that?
The most obvious reason is the odd hours throw a person’s 24-hour biological clock—and metabolism—out of sync. But an NIH-funded team of researchers has new evidence in mice to suggest the answer could go deeper to include the trillions of microbes that live in our guts—and, more specifically, the way they “talk” to intestinal cells. Their studies suggest that what gut microbes “say” influences the activity of a key clock-driven protein called NFIL3, which can set intestinal cells up to absorb and store more fat from the diet while operating at hours that might run counter to our fixed biological clocks.
Share this:
- Click to share on LinkedIn (Opens in new window)
- Click to share on Pinterest (Opens in new window)
- Click to share on Tumblr (Opens in new window)
- Click to share on Reddit (Opens in new window)
- Click to share on Telegram (Opens in new window)
- Click to share on WhatsApp (Opens in new window)
- Click to print (Opens in new window)
Cardiometabolic Disease: Big Data Tackles a Big Health Problem
Posted on by Dr. Francis Collins
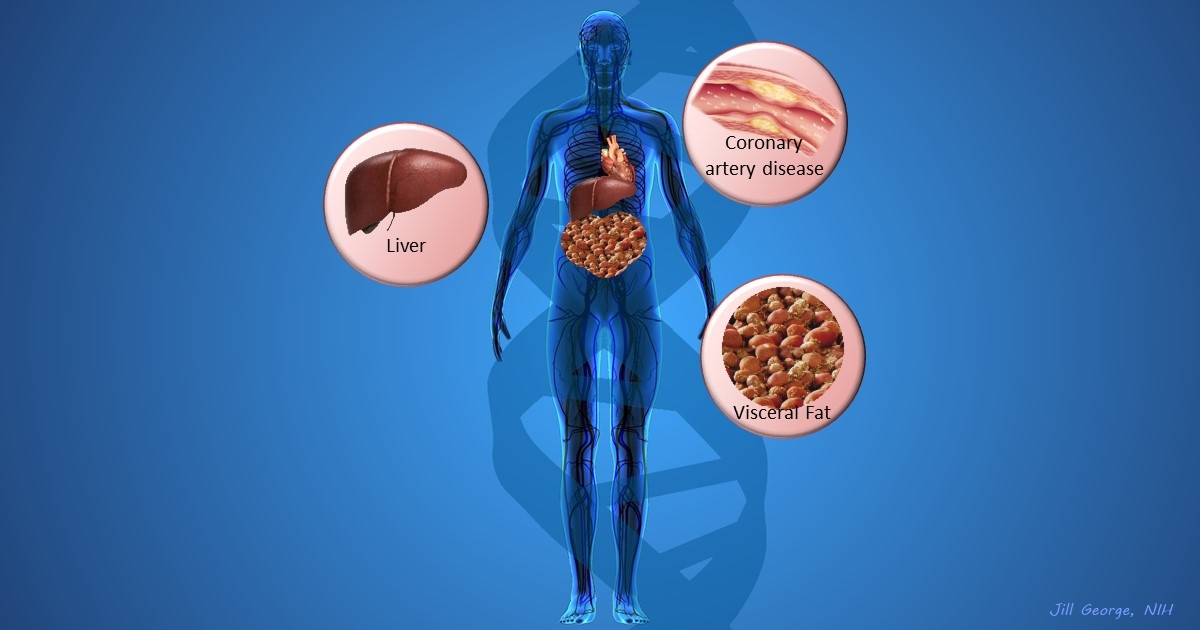
More and more studies are popping up that demonstrate the power of Big Data analyses to get at the underlying molecular pathology of some of our most common diseases. A great example, which may have flown a bit under the radar during the summer holidays, involves cardiometabolic disease. It’s an umbrella term for common vascular and metabolic conditions, including hypertension, impaired glucose and lipid metabolism, excess belly fat, and inflammation. All of these components of cardiometabolic disease can increase a person’s risk for a heart attack or stroke.
In the study, an international research team tapped into the power of genomic data to develop clearer pictures of the complex biocircuitry in seven types of vascular and metabolic tissue known to be affected by cardiometabolic disease: the liver, the heart’s aortic root, visceral abdominal fat, subcutaneous fat, internal mammary artery, skeletal muscle, and blood. The researchers found that while some circuits might regulate the level of gene expression in just one tissue, that’s often not the case. In fact, the researchers’ computational models show that such genetic circuitry can be organized into super networks that work together to influence how multiple tissues carry out fundamental life processes, such as metabolizing glucose or regulating lipid levels. When these networks are perturbed, perhaps by things like inherited variants that affect gene expression, or environmental influences such as a high-carb diet, sedentary lifestyle, the aging process, or infectious disease, the researchers’ modeling work suggests that multiple tissues can be affected, resulting in chronic, systemic disorders including cardiometabolic disease.
Share this:
- Click to share on LinkedIn (Opens in new window)
- Click to share on Pinterest (Opens in new window)
- Click to share on Tumblr (Opens in new window)
- Click to share on Reddit (Opens in new window)
- Click to share on Telegram (Opens in new window)
- Click to share on WhatsApp (Opens in new window)
- Click to print (Opens in new window)
Tags: bad cholesterol, big data, bioinformatics, blood lipids, cardiometabolic disease, cardiovascular disease, coronary artery disease, coronary bypass surgery, drug delivery, eQTL, Estonia, fat, gemomics, gene networks, gene variants, GWAS, hyperlipidemia, hypertension, LDL, liver, metabolic syndrome, NHGRI GWAS Catalog, PCSK9, PMI, Precision Medicine Initiative Cohort Program, STARNET, Sweden, systems biology, systems genetics, type 2 diabetes, visceral abdominal fat, visceral fat
