computational genomics
Crowdsourcing 600 Years of Human History
Posted on by Dr. Francis Collins
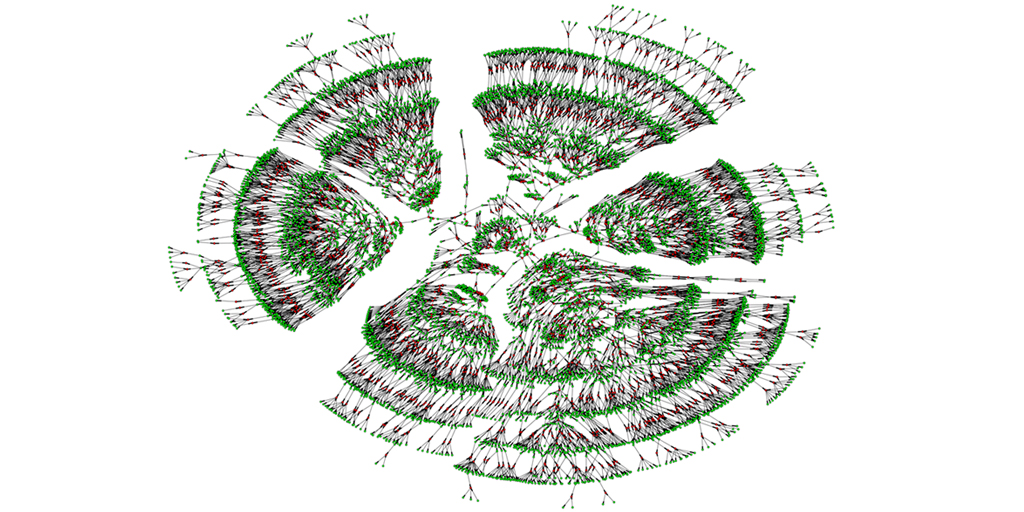
Caption: A 6,000-person family tree, showing individuals spanning seven generations (green) and their marital links (red).
Credit: Columbia University, New York City
You may have worked on constructing your family tree, perhaps listing your ancestry back to your great-grandparents. Or with so many public records now available online, you may have even uncovered enough information to discover some unexpected long-lost relatives. Or maybe you’ve even submitted a DNA sample to one of the commercial sources to see what you could learn about your ancestry. But just how big can a family tree grow using today’s genealogical tools?
A recent paper offers a truly eye-opening answer. With permission to download the publicly available, online profiles of 86 million genealogy hobbyists, most of European descent, the researchers assembled more than 5 million family trees. The largest totaled more than 13 million people! By merging each tree from the crowd-sourced and public data, including the relatively modest 6,000-person seedling shown above, the researchers were able to go back 11 generations on average to the 15th century and the days of Christopher Columbus. Doubly exciting, these large datasets offer a powerful new resource to study human health, having already provided some novel insights into our family structures, genes, and longevity.
Creative Minds: Using Machine Learning to Understand Genome Function
Posted on by Dr. Francis Collins
Science has always fascinated Anshul Kundaje, whether it was biology, physics, or chemistry. When he left his home country of India to pursue graduate studies in electrical engineering at Columbia University, New York, his plan was to focus on telecommunications and computer networks. But a course in computational genomics during his first semester showed him he could follow his interest in computing without giving up his love for biology.
Now an assistant professor of genetics and computer science at Stanford University, Palo Alto, CA, Kundaje has received a 2016 NIH Director’s New Innovator Award to explore not just how the human genome sequence encodes function, but also why it functions in the way that it does. Kundaje even envisions a time when it might be possible to use sophisticated computational approaches to predict the genomic basis of many human diseases.

