Of Microbes, Molecules, and Maps
Posted on by Dr. Francis Collins
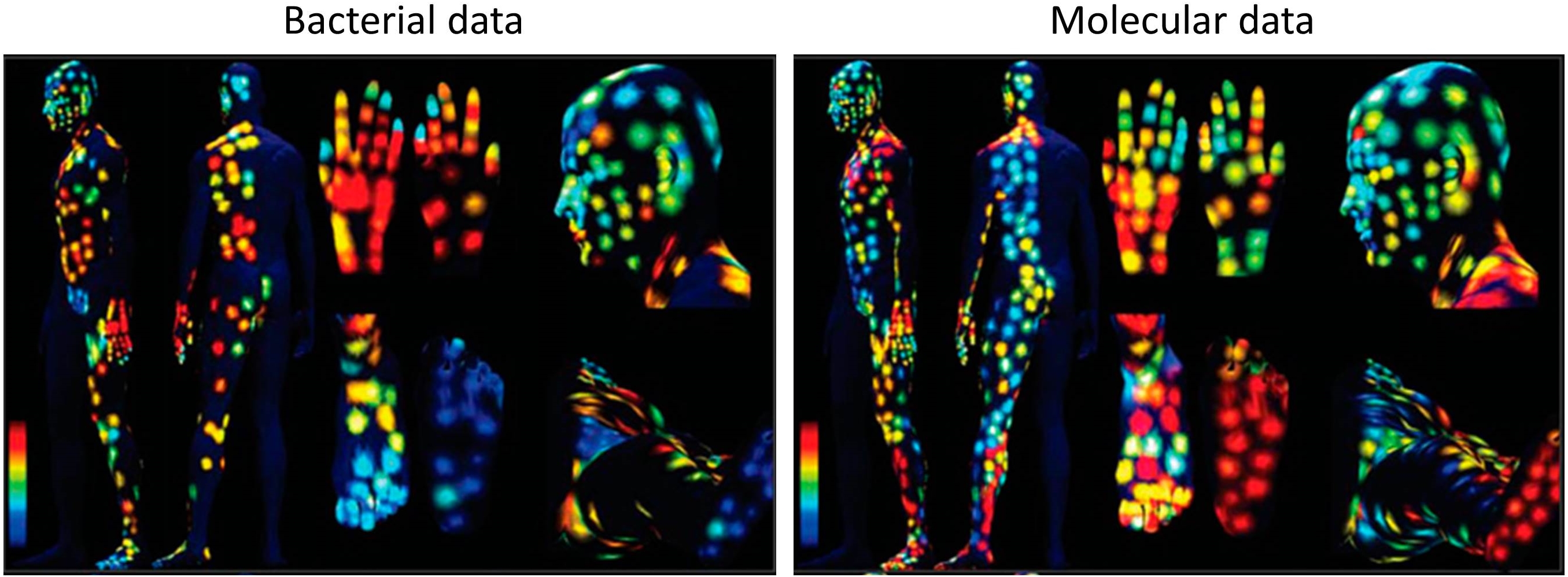
These glow-in-the-dark images may look like a 60’s rock album cover, but they’re actually a reflection of some way cool science. Here are maps showing the diversity of bacteria (left) and “acquired” molecules (right) on the skin of a healthy man. Blue indicates areas of least diversity; green/yellow, medium; and orange/red, the greatest.
To create these maps, NIH-funded researchers at the University of California, San Diego (UCSD), and their colleagues swabbed the skin of a male volunteer at roughly 400 spots to sample for bacteria. Then, they swabbed the same spots again to sample for chemicals and other types of molecules, natural or synthetic, that the man’s skin acquired over the course of his daily activities. Examples of such molecules include chemicals in shampoo and grooming products, polymers shed from clothing, and proteins released when skin cells are damaged or die.
Using ribosomal RNA sequencing, mass spectroscopy, and other sophisticated laboratory techniques, the researchers then identified the bacteria and molecules contained in each sample and quantified their diversity at that specific site. The data were then analyzed and translated into the rainbow-hued topographical maps that you see above.
These and a collection of similar maps, just published in the journal PNAS [1], represent a technical advance in the effort to take distinctly different types of Big Data and visualize them in a unified format. Pieter Dorrestein, who co-authored the study with Rob Knight, Theodore Alexandrov, and colleagues, says the secret was to use a consistent spatial orientation to organize diverse categories of scientific information. This shared orientation enables a viewer to compare maps of different data types more easily. For example, the new mapping system will enable researchers to study with greater precision how the skin’s molecular environment affects the microbial communities that live on it, possibly providing clues to such common skin conditions as allergic dermatitis and psoriasis.
To give you an idea of how this might work, try comparing these two maps of a healthy woman volunteer’s body. The image on the left shows the density and distribution of Propionibacterium, a common genus of bacteria. The image on the right shows the density and distribution of oleic acid, a fatty acid found in human cells and also used in soap, shampoos, lotions, and other cosmetics. High densities are indicated red; medium, yellow/green; low, blue; and undetectable, black.
As you can see, Propionibacterium tends to colonize the head, face, neck, and many other areas of the body where the molecular map shows oleic acid—locations that just happen to be where soap, shampoo, lotions, and other cosmetics are often applied. This kind of data can generate lots of hypotheses. The microbiome is really coming into its own!
Reference:
3D molecular cartography of the human skin surface. Bouslimani A, Porto C, Rath CM, Knight R, Alexandrov T, Dorrestein PC, et. al. Proc Natl Acad Sci USA. 2015 March 30. Pii: 201424409. [Epub ahead of print]
Links:
Skin Conditions (Medline Plus/NIH)
Dorrestein Lab, University of California, San Diego
NIH Big Data to Knowledge (BD2K/NIH)
Human Microbiome Project (Common Fund/NIH)
NIH Support: National Institute of General Medical Sciences


You are doing great work, thank you.
So interesting, thanks
What is the resolving power of these images, i.e., How well defined are the boundaries?